Screen It™ CRISPR Cas9 Cleavage Detection Kit
|
Cat. No.
|
G990
Print
|
|
Name
|
Screen It™ CRISPR Cas9 Cleavage Detection Kit
|
|
Unit
|
100 Reactions
|
|
Category
|
Cas Proteins & CRISPR Screening
|
|
Description
|
abm’s Screen It™ CRISPR Cas9 Cleavage Detection Kit is a robust and precise RNA-dependent assay designed to detect successful insertions and deletions (indels) in genomic DNA following CRISPR-Cas9 editing. The kit can determine whether a clone is monoallelic (mutations in one copy), biallelic (mutations in both copies), or wild-type (unchanged), offering a key screening advantage over the standard T7E1 Surveyor Assay.
The assay begins by amplifying the sgRNA target site from CRISPR-edited cells using PCR. The resulting amplicon is then cleaved by a ribonucleoprotein (RNP) complex, generating a distinct banding pattern that is easily resolved on an agarose gel. This clear visual output allows users to quickly and confidently assess editing outcomes.
By streamlining the post-editing screening process, the Screen It™ Kit helps researchers save time and effort, providing reliable genotyping results for efficient identification of successfully edited clones.
| Product Component |
Quantity |
| Cell Lysis Buffer |
1.25 ml |
| Protein Degrader |
100 µl |
| Scaffold Template and Primer Mix |
100 µl |
| 2X sgRNA Synthesis Buffer |
100 µl |
| sgRNA Synthesis Enzyme Mix |
50 µl |
| sgRNA Control Oligo |
20 µl |
| Wild Type Control Primer and Template Mix |
20 µl |
| Monoallelic Control Primer and Template Mix |
20 µl |
| Biallelic Control Primer and Template Mix |
20 µl |
| RNP Degrader |
100 µl |
| spCas9 Nuclease Protein |
250 µl |
| 10X Cas9 Reaction Buffer |
1.25 ml |
| MegaFi™ Pro Fidelity 2X MasterMix |
2 x 1.25 ml |
Additional Materials Required
- Target-specific Primers
(abm offers sgRNA PCR Primer Pair Design & Synthesis Service, Cat. No. C336)
- sgRNA Target-specific Oligo
(abm offers sgRNA Target-Specific Oligo Design & Synthesis Service, Cat. No. C337)
- CRISPR-edited Cells
- Wild-type Cells
- Nuclease-free H2O
|
|
Storage Condition
|
Store all components at -20°C.
|
|
Material Citation
|
If use of this material results in a scientific publication, please cite the material in the following manner: Applied Biological Materials Inc, Cat. No. G990
|
|
|
What is the Screen It™ CRISPR Cas9 Cleavage Detection Kit used for?
|
|
This kit is designed to identify successful insertions or deletions (indels) in genomic DNA following CRISPR/Cas9 editing. It can determine if gene editing is monoallelic, biallelic, or unedited, providing valuable insight into the efficiency of the CRISPR process and the editing pattern.
|
|
|
How does the Screen It™ kit work?
|
|
The Screen It™ CRISPR Cas9 Cleavage Detection Kit follows a streamlined process to identify successful CRISPR-induced insertions or deletions:
- Cell Lysis: CRISPR-edited and wild-type cells are lysed to extract genomic DNA, which is prepared for PCR amplification.
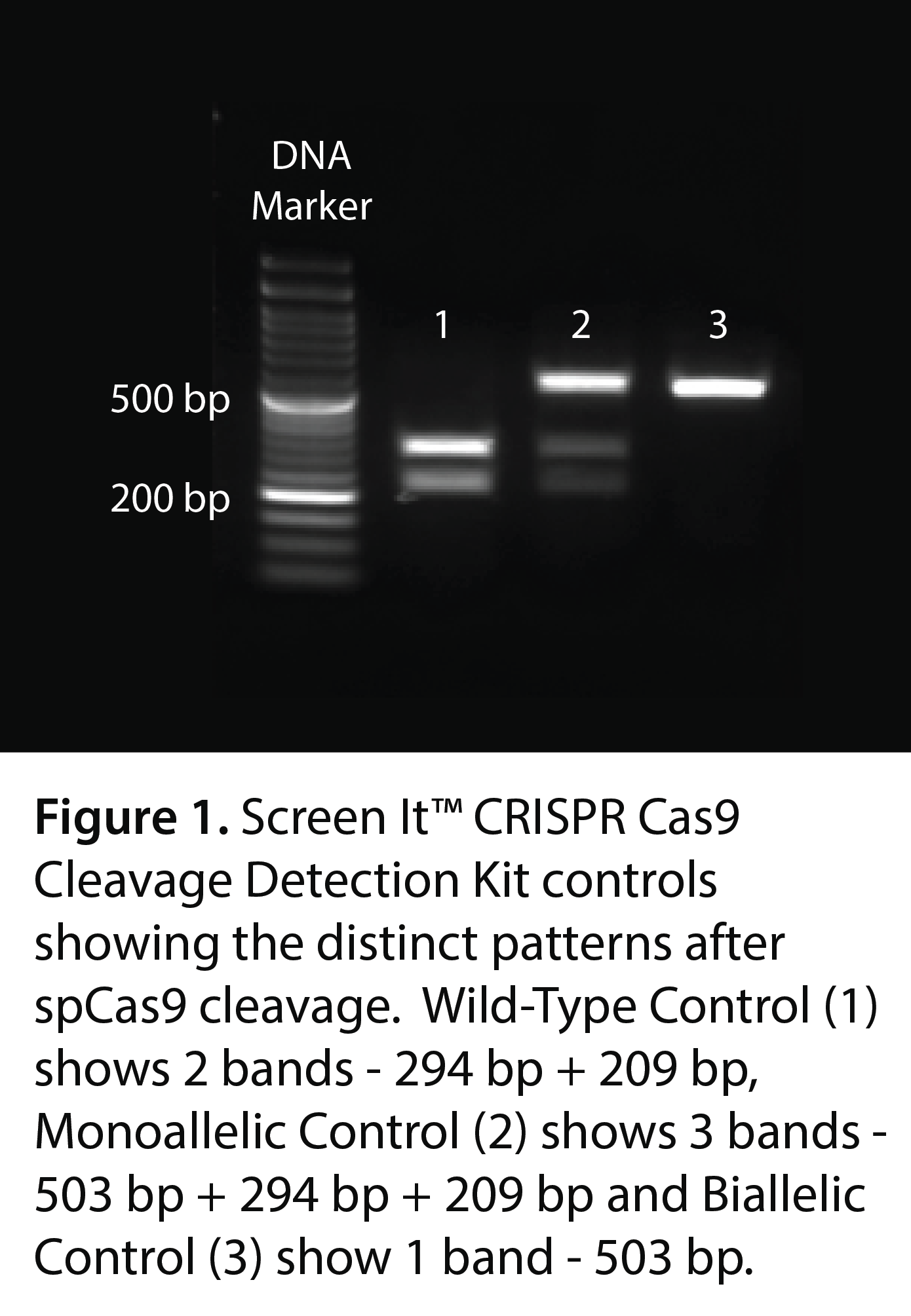
- PCR Amplification: The sgRNA target region is amplified using specific primers. The PCR products are analyzed on a gel, with control amplicons expected around 503 bp.
- sgRNA Synthesis: sgRNA is synthesized using the provided template and enzyme mix for cleavage in the next step.
- In Vitro Cas9 Cleavage: The sgRNA and Cas9 protein form an RNP complex that cleaves the PCR products, generating distinct bands based on the editing outcome.
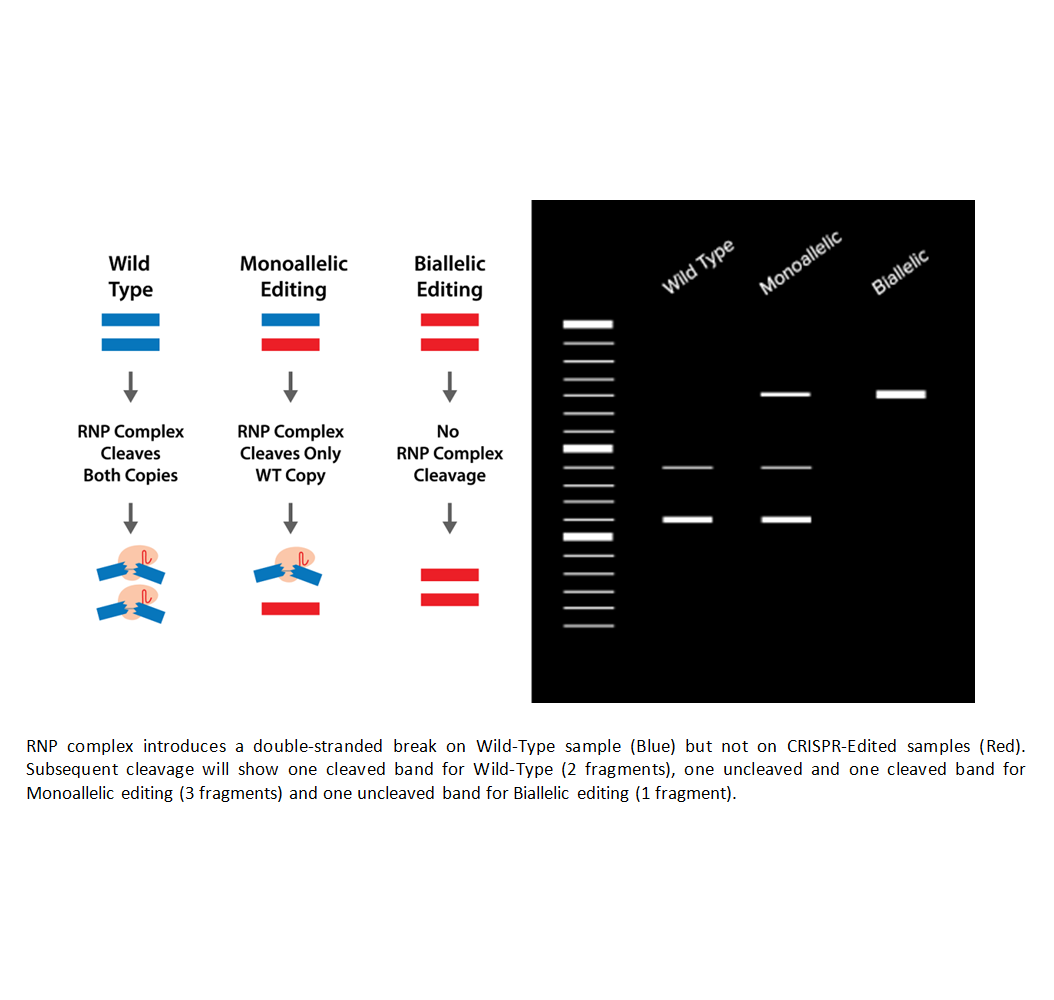
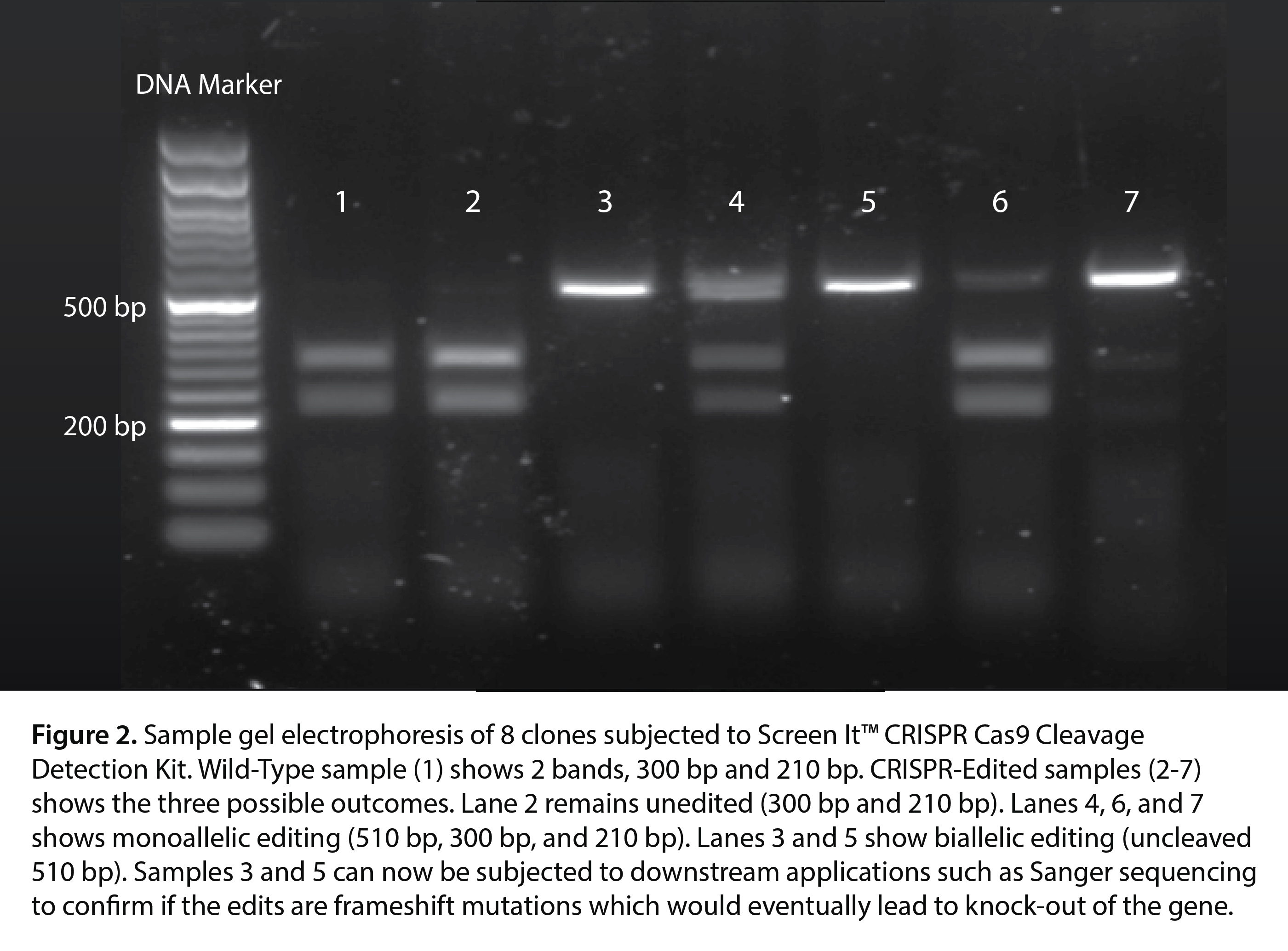
- Gel Analysis: The cleavage patterns are analyzed on an agarose gel, where wild-type cells show two bands, biallelic edits show one band, and monoallelic edits show three bands.
This kit helps efficiently determine the editing status (monoallelic, biallelic, or wild-type) of your CRISPR-edited samples.
|
|
|
How do I prepare the PCR primers for this kit?
|
|
Primers should amplify the sgRNA target region and the Cas9 cut site, ensuring the cut site is not in the middle of the amplicon for clear cleavage patterns. If needed, abm offers a primer design service ( Cat. No. C336) to help ensure that primers are optimized for your specific target and CRISPR experiment.
|
|
|
How do I analyze the cleavage patterns on a gel?
|
|
To analyze the cleavage patterns on a gel:
- Wild-type samples will show two bands, indicating that the Cas9 enzyme cleaved the DNA at the target site.
- Monoallelic editing (one copy edited) will show three bands: one for the unedited wild-type allele and two for the edited alleles.
- Biallelic editing (both copies edited) will show one band, representing the edited alleles.
These band patterns help you determine if the cells are wild-type, monoallelic, or biallelic after CRISPR editing.
|
|
|
What do I need to synthesize sgRNA for use with this kit?
|
|
The kit includes a scaffold template and primer mix to transcribe sgRNA. Users need to input their sgRNA sequence into the kit for synthesis, which will be used to generate sgRNA suitable for the RNP cleavage reactions in the assay.
|
|
|
Can I use this kit for high-throughput applications?
|
|
Yes, the kit’s streamlined protocol is designed for ease of use and rapid processing, making it suitable for high-throughput applications in genomic screening. It can help researchers quickly identify edited clones and save valuable time in CRISPR experiment workflows.
|
There are no references for this product yet!
This product has no review yet.
Controls and Related Product:
UNIT
40µg (250pmol/ 25µL)
UNIT
32.5µg (250pmol/ 25µL)
{"id":3236,"cat_no":"G990","sku_in_m2":"","old_cat_no":"","name":"Screen It\u2122 CRISPR Cas9 Cleavage Detection Kit","category_id":208,"category_name":"Cas Proteins \u0026 CRISPR Screening","frontend_category_id":71,"frontend_category_ids":"57,67,71","gene_id":0,"gene_name":"","gene_full_name":"","alias":"","unit_quantity":"100 Reactions","accession_number":"","species_id":0,"species":"","raised_in_id":0,"raised_in":"","cell_type_id":0,"cell_type":"","tissue_id":0,"tissue":"","price":"705.0000","price_before_discount":"1500.0000","keywords":"","invisible_search_keywords":"CRISPR Cas9 Cleavage Detection Kit, crispr cleavage detection kit, rna-dependent assay, crispr cas9, monoallelic, biallelic, wildtype, Genomic Cleavage Detection Kit, Genome Editing Detection Kit, genome editing, Cas9 screening, CRISPR Cas9 screening, CRISPR Screening kit, surveyor assay, crispr test assay","filter_group_id":34,"filter_group_name":"Molecular Biology","filter_id":41,"filter_name":"Kits \u0026 Other Enzymes","search_filter_ids":"32,34","search_filter_keys":"32-34","temp_field":null,"customization":0,"filter_table_id":0,"filter_table_name":"","related_group_id":17,"url":"","operator":"","points_rule":1,"status":1,"need_remove":0,"document_sort":"","price_old":"1500.0000","price2024":"1500.0000","created_at":"2023-03-29 22:43:03","updated_at":"2025-03-24 20:46:26","Confirmed":"Need Confirm","geo_price_before_discount":"1500.00","geo_titer7_price_before_discount":"1649.00","cate_url":"cas9-proteins.html","frontendBreadcrumbs":[{"id":57,"pid":86,"name":"Genetic Materials","page_id":8,"page_type":"App\\Models\\Category","href":null,"sort":57,"show_in_nav":1,"show_in_homepage":1,"has_children":1,"status":1,"related_table":null,"created_at":null,"updated_at":"2025-01-20 08:46:24","url_key":"genetic-materials.html","url":"genetic-materials.html"},{"id":67,"pid":57,"name":"CRISPR","page_id":15,"page_type":"App\\Models\\Category","href":null,"sort":67,"show_in_nav":1,"show_in_homepage":1,"has_children":1,"status":1,"related_table":null,"created_at":null,"updated_at":null,"url_key":"CRISPR-Cas9-sgRNA.html","url":"CRISPR-Cas9-sgRNA.html"},{"id":71,"pid":67,"name":"Cas Proteins \u0026 CRISPR Screening","page_id":223,"page_type":"App\\Models\\Cms","href":null,"sort":71,"show_in_nav":1,"show_in_homepage":1,"has_children":0,"status":1,"related_table":null,"created_at":null,"updated_at":null,"url_key":"cas9-proteins.html","url":"cas9-proteins.html"}],"price_flag":0,"builderURL":"","builderViewerURL":"","mapURL":"","viewerURL":"","builderType":"","productType":"App\\Models\\CatalogBaseMolecular","geo_price":"705.00","geo_price_symbol":"$705.00","category":{"id":208,"categories":"Cas Proteins \u0026 CRISPR Screening","pid":15,"url_key":"cas9-proteins.html","css":null,"wid":0,"breadcrumbs":null,"breadcrumbsIdJson":"","breadcrumbsHtml":"","description":"\u003Cp\u003ECas9 proteins\u003C\/p\u003E\n\u003Cdiv id=\u0022gtx-trans\u0022 style=\u0022position: absolute; left: -35px; top: -10.5px;\u0022\u003E\n\u003Cdiv class=\u0022gtx-trans-icon\u0022\u003E\u003C\/div\u003E\n\u003C\/div\u003E","meta_title":"Cas Proteins \u0026 CRISPR Screening","meta_keywords":"Cas Proteins \u0026 CRISPR Screening","meta_description":"Cas Proteins \u0026 CRISPR Screening","deleted_at":null,"enable":"N","parent_list":"8,15","table_name":null,"image":null,"independentPage":0,"top_type":1,"sort_order":null,"in_footer":1,"fid":null,"created_at":"2024-08-28 23:24:53","updated_at":"2024-12-19 17:58:36"},"info":{"id":18444,"cat_no_base":null,"parent_id":3236,"description":"\u003Cp\u003E\u003Cstrong\u003Eabm\u003C\/strong\u003E\u0026rsquo;s \u003Cstrong\u003EScreen It\u0026trade; CRISPR Cas9 Cleavage Detection Kit\u003C\/strong\u003E is a \u003Cstrong\u003Erobust and precise RNA-dependent assay\u003C\/strong\u003E designed to detect successful insertions and deletions (indels) in genomic DNA following CRISPR-Cas9 editing. The kit can determine whether a clone is monoallelic (mutations in one copy), biallelic (mutations in both copies), or wild-type (unchanged), offering \u003Cstrong\u003Ea key screening advantage over the standard T7E1 Surveyor Assay\u003C\/strong\u003E.\u003C\/p\u003E\n\u003Cp\u003EThe assay begins by amplifying the sgRNA target site from CRISPR-edited cells using PCR. The resulting amplicon is then cleaved by a ribonucleoprotein (RNP) complex, generating a distinct banding pattern that is easily resolved on an agarose gel. This clear visual output allows users to quickly and confidently assess editing outcomes.\u003C\/p\u003E\n\u003Cp\u003EBy streamlining the post-editing screening process, the Screen It\u0026trade; Kit helps researchers save time and effort, providing reliable genotyping results for efficient identification of successfully edited clones.\u003C\/p\u003E\n\u003Ctable class=\u0022bootstrap-table bootstrap-table-hover abm-service-table abm-perfect-table2\u0022 style=\u0022margin: 20px 0;\u0022\u003E\n\u003Cthead\u003E\n\u003Ctr\u003E\n\u003Cth\u003E\u003Cstrong\u003EProduct Component\u003C\/strong\u003E\u003C\/th\u003E\n\u003Cth\u003E\u003Cstrong\u003EQuantity\u003C\/strong\u003E\u003C\/th\u003E\n\u003C\/tr\u003E\n\u003C\/thead\u003E\n\u003Ctfoot\u003E\n\u003Ctr\u003E\n\u003Ctd\u003ECell Lysis Buffer\u003C\/td\u003E\n\u003Ctd\u003E1.25 ml\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EProtein Degrader\u003C\/td\u003E\n\u003Ctd\u003E100 \u0026micro;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EScaffold Template and Primer Mix\u003C\/td\u003E\n\u003Ctd\u003E100 \u0026micro;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003E2X sgRNA Synthesis Buffer\u003C\/td\u003E\n\u003Ctd\u003E100 \u0026micro;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EsgRNA Synthesis Enzyme Mix\u003C\/td\u003E\n\u003Ctd\u003E50 \u0026micro;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EsgRNA Control Oligo\u003C\/td\u003E\n\u003Ctd\u003E20 \u0026micro;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EWild Type Control Primer and Template Mix\u003C\/td\u003E\n\u003Ctd\u003E20 \u0026micro;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EMonoallelic Control Primer and Template Mix\u003C\/td\u003E\n\u003Ctd\u003E20 \u0026micro;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EBiallelic Control Primer and Template Mix\u003C\/td\u003E\n\u003Ctd\u003E20 \u0026micro;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003ERNP Degrader\u003C\/td\u003E\n\u003Ctd\u003E100 \u0026micro;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EspCas9 Nuclease Protein\u003C\/td\u003E\n\u003Ctd\u003E250 \u0026micro;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003E10X Cas9 Reaction Buffer\u003C\/td\u003E\n\u003Ctd\u003E1.25 ml\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EMegaFi\u0026trade; Pro Fidelity 2X MasterMix\u003C\/td\u003E\n\u003Ctd\u003E2 x 1.25 ml\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003C\/tfoot\u003E\n\u003C\/table\u003E\n\u003Cp\u003E\u003Cstrong\u003EAdditional Materials Required\u003C\/strong\u003E\u003C\/p\u003E\n\u003Cul\u003E\n\u003Cli\u003ETarget-specific Primers \u003Cbr \/\u003E(\u003Cstrong\u003Eabm\u003C\/strong\u003E offers sgRNA PCR Primer Pair Design \u0026 Synthesis Service, \u003Ca href=\u0022\/Primer-Design-Synthesis.html\u0022\u003ECat. No. C336\u003C\/a\u003E)\u003C\/li\u003E\n\u003Cli\u003EsgRNA Target-specific Oligo \u003Cbr \/\u003E(\u003Cstrong\u003Eabm\u003C\/strong\u003E offers sgRNA Target-Specific Oligo Design \u0026 Synthesis Service, \u003Ca href=\u0022\/Primer-Design-Synthesis.html\u0022\u003ECat. No. C337\u003C\/a\u003E)\u003C\/li\u003E\n\u003Cli\u003ECRISPR-edited Cells\u003C\/li\u003E\n\u003Cli\u003EWild-type Cells\u003C\/li\u003E\n\u003Cli\u003ENuclease-free H\u003Csub\u003E2\u003C\/sub\u003EO\u003C\/li\u003E\n\u003C\/ul\u003E","disclaimer":null,"application":null,"components":null,"cas9_origin":null,"concentration":null,"enzymes_size":null,"genecraft_series":null,"guarantee":null,"population":null,"qc":null,"format_general":null,"including_screening_kit":null,"expression_system_general":null,"purity":null,"image":null,"insert_size":null,"shipping_conditions":null,"source_catalog_number":null,"inactivation_protocol":null,"led_viewer_compatibility":null,"unit_definition":null,"vector":null,"reaction_buffer":null,"storage_buffer":null,"caution":null,"storage_condition":"Store all components at -20\u00b0C.","product_volume":null,"reporter":null,"safeview_series":null,"source_catno":null,"stain_color":null,"supplier":null,"internal_supplier":null,"internal_note":null,"inventory_location":"PCR-2A","note":null,"recommend":null,"depositor":null,"licensor_name":null,"licensor_contact_information":null,"contract_termination_date":null,"royalty_rates":null,"cas_type":null,"cas_origin":null,"cas_protein_marker":null,"source":null,"endotoxin_level":null,"additional_information":null,"titer":null,"nucleotide_format":null,"protocol_overview":null,"source_price":null,"created_at":"2023-03-29 22:43:03","updated_at":"2025-03-24 20:46:26","short_description":null,"reaction_definition":null,"specificity":null,"promotions":null},"maps":[],"media":[{"id":382620,"parent_id":3236,"parent_type":"App\\Models\\CatalogBaseMolecular","file_path":"\/upload\/PeBdgoZ3xchpb3NW2JCmFfpo78Th0oxzTUZ9X3Zp.png","title":null,"text":null,"file_type":"image","alt":null,"url":null,"position":null,"status":1,"entity_id_m2":null,"sku_in_m2":null,"value_id_m2":null,"attribute_id":0,"created_at":"2023-06-12 17:20:10","updated_at":"2023-06-12 17:20:10"},{"id":382621,"parent_id":3236,"parent_type":"App\\Models\\CatalogBaseMolecular","file_path":"\/upload\/iMKV5Qc6MRgEzwvkeN9EdxWibsvBI2Zm00oxwdia.png","title":null,"text":null,"file_type":"image","alt":null,"url":null,"position":null,"status":1,"entity_id_m2":null,"sku_in_m2":null,"value_id_m2":null,"attribute_id":0,"created_at":"2023-06-12 17:20:16","updated_at":"2023-06-12 17:20:16"},{"id":382622,"parent_id":3236,"parent_type":"App\\Models\\CatalogBaseMolecular","file_path":"\/upload\/L4fsqn117piJ2iiefmvKFsfzR5zfUEfyxw1ehrrW.png","title":null,"text":null,"file_type":"image","alt":null,"url":null,"position":null,"status":1,"entity_id_m2":null,"sku_in_m2":null,"value_id_m2":null,"attribute_id":0,"created_at":"2023-06-12 17:20:21","updated_at":"2023-06-12 17:20:21"}],"gene":null}