CRISPR Genomic Cleavage Detection Kit
|
Cat. No.
|
G932
Print
|
|
Name
|
CRISPR Genomic Cleavage Detection Kit
|
|
Unit
|
100 Reactions
|
|
Category
|
Cas Proteins & CRISPR Screening
|
|
Description
|
abm’s CRISPR Genomic Cleavage Detection Kit is a simple and rapid assay designed to verify your genomic editing process. Precision is essential in genome editing and gene modification, and confirming successful gene editing early can save both time and resources. This kit allows you to easily detect genomic cleavage, ensuring accurate results in your experiments.
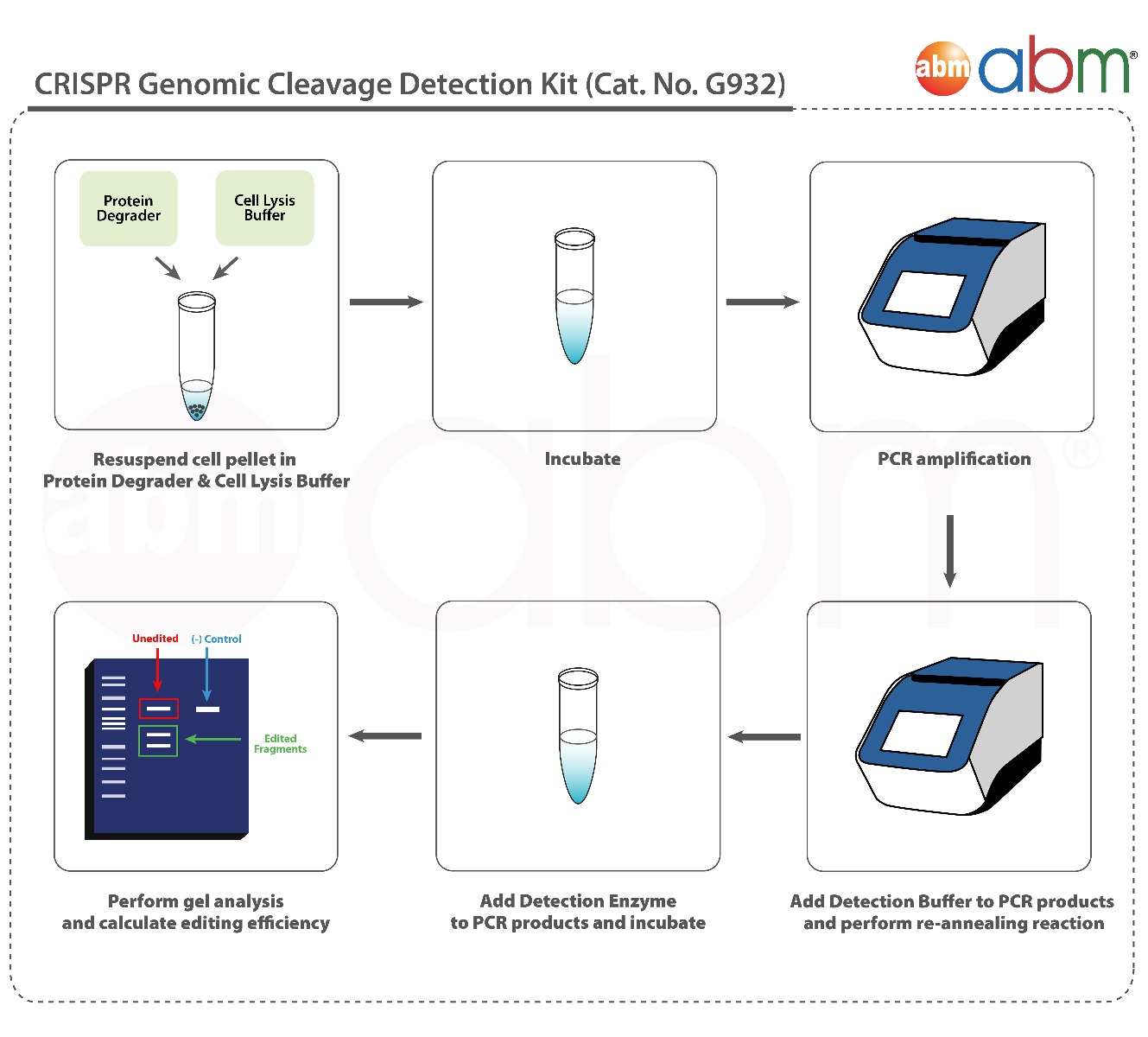
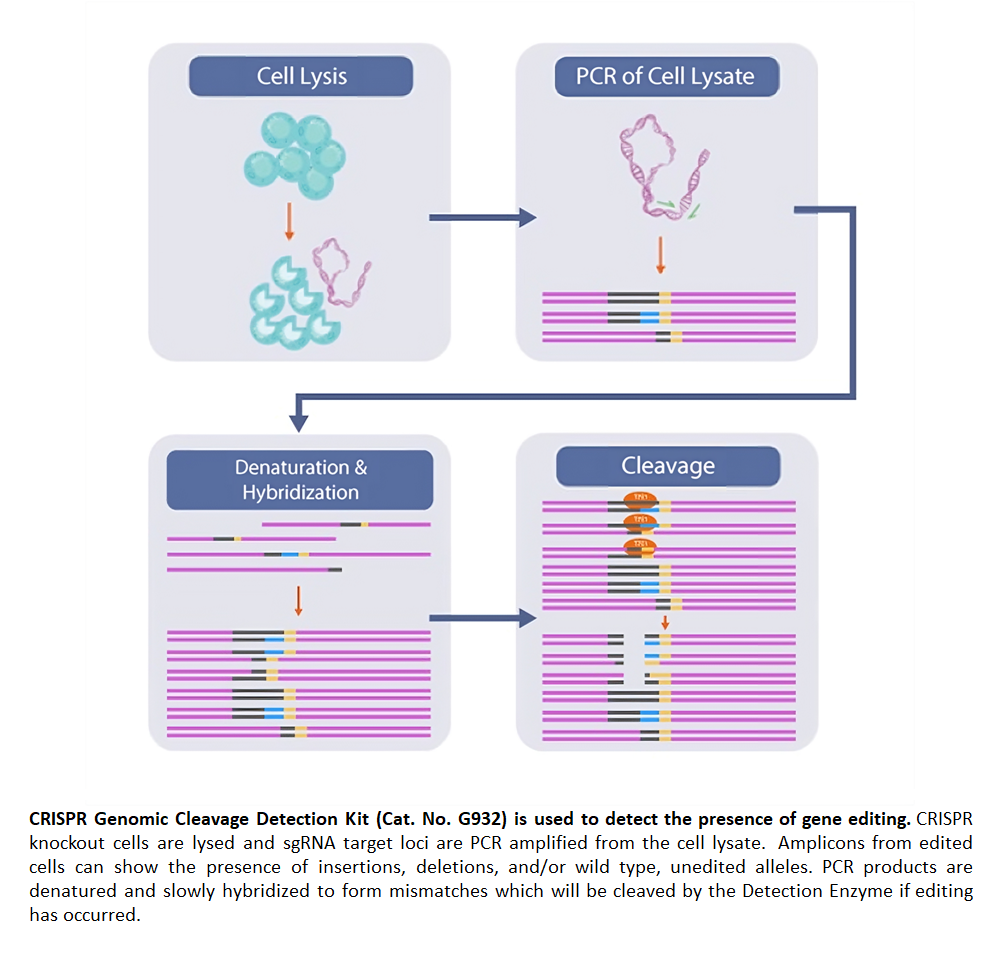
The kit uses CRISPR-edited samples as templates in PCR reactions targeting your specific region of interest. It requires a pair of primers flanking each sgRNA target site to detect genomic cleavage, which can be ordered from abm (Cat. No. C336 - sgRNA PCR Primer Pair Design & Synthesis Service). After PCR amplification, the products are denatured and re-annealed, creating mismatches within the double-strand DNA. Our detection enzyme recognizes these mismatches and cleaves the strands to produce distinguishable band sizes upon gel analysis.
The ready-to-use kit includes all necessary reagents, including a control template and primers to ensure reliable results. The control template produces bands of approximately 750bp after PCR amplification and 500bp and 250bp after enzymatic cleavage. With a quick processing time, this assay is a valuable addition to any genomic-editing workflow.
Kit Features:
- Ease of use with simple steps
- Rapid set-up
- Streamlined protocol suitable for high-throughput applications
| Product Component |
Quantity |
| Cell Lysis Buffer |
1.25 ml |
| Protein Degarder |
100µl |
| Detection Enzyme |
50 μl |
| 10X Detection Buffer |
200 μl |
| Control Primer & Template |
10 μl |
| MegaFi™ Pro Fidelity 2X MasterMix |
2 x 1.25 ml |
|
|
Storage Condition
|
Store all components at -20°C.
|
|
Material Citation
|
If use of this material results in a scientific publication, please cite the material in the following manner: Applied Biological Materials Inc, Cat. No. G932
|
|
|
How does this kit verify gene editing?
|
|
The CRISPR Genomic Cleavage Detection Kit is designed to verify CRISPR-based genome editing by detecting mismatches in PCR products. After PCR amplification of CRISPR-edited samples, the products are denatured and re-annealed, creating mismatches at the edited site. The detection enzyme cleaves these mismatches, producing distinguishable fragments that can be analyzed on a gel. The kit ensures precision by confirming successful edits early in the process, saving time and resources.
|
|
|
What should I expect to see on the gel when using the CRISPR Genomic Cleavage Detection Kit?
|
|
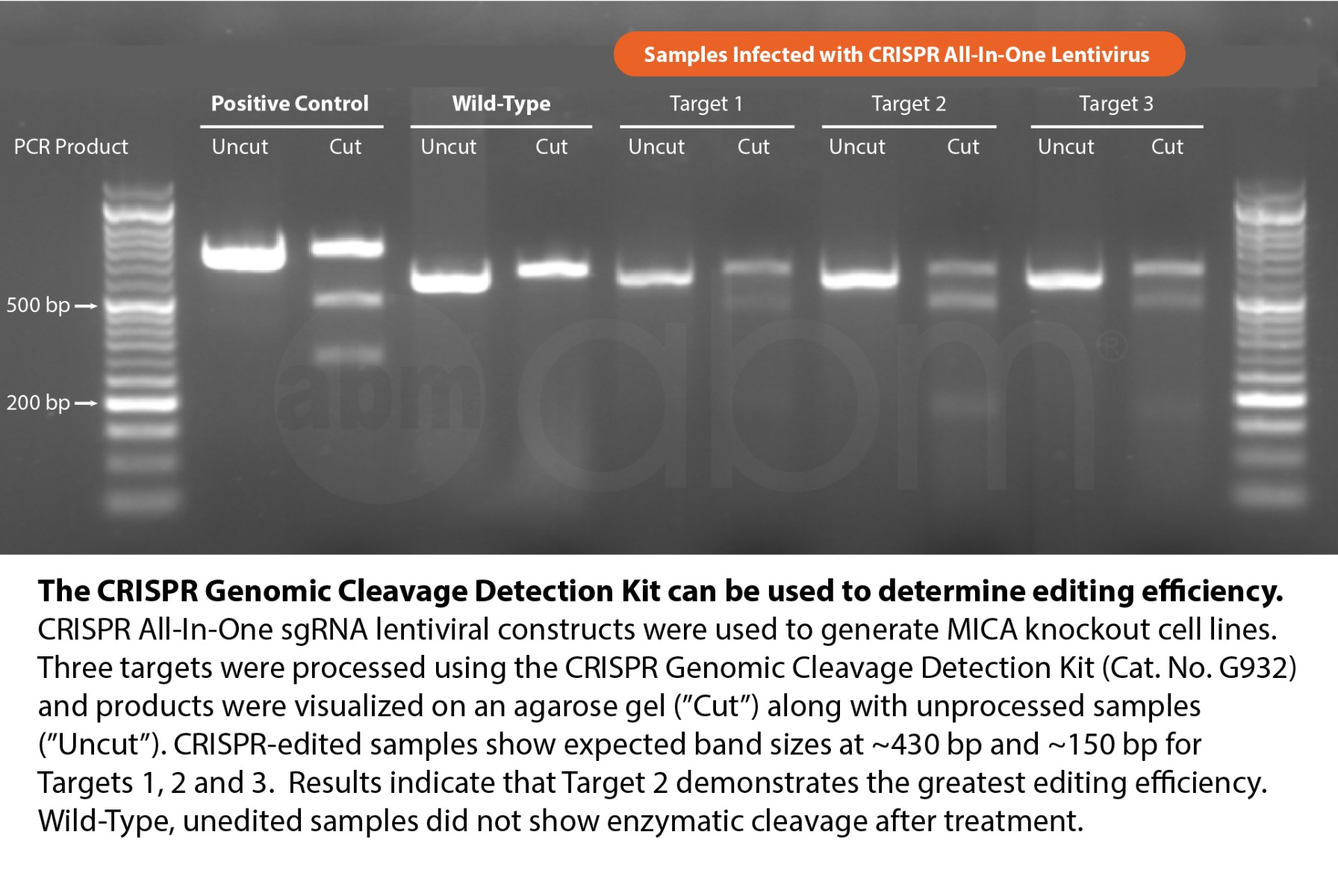
On the gel, you should expect the following:
- Control Template (Wild-Type Cells)
- Before Cleavage: A 750 bp band.
- After Cleavage: Two bands at 500 bp and 250 bp.
- CRISPR-Edited Cells
- Before Cleavage: A 750 bp band.
- After Cleavage: Two bands at 500 bp and 250 bp (if editing was successful). If editing was unsuccessful, the 750 bp band may remain.
The presence of 500 bp and 250 bp bands indicates successful CRISPR editing.
|
|
|
What is the control template used for in this kit?
|
|
The control template serves as a reference to verify the performance of the assay. It produces PCR products of approximately 750 bp, and after cleavage, it produces bands of 500 bp and 250 bp. These control bands help confirm the assay’s accuracy.
|
-
Chakraborty, A., Dutta, A., Dettori, L. G., Daoud, R., Li, J., Gonzalez, L., ... & Feng, W. (2024). Complex interplay between FMRP and DHX9 during DNA replication stress. Journal of Biological Chemistry, 300(1). https://doi.org/10.1016/j.jbc.2023.105572
Chakraborty, A., Dutta, A., Dettori, L. G., Li, J., Gonzalez, L., Xue, X., ... & Feng, W. (2021). FMRP directly interacts with R-loop and shows complex interplay with the DHX9 helicase. bioRxiv, 2021-04. https://doi.org/10.1101/2021.04.21.440759
Ortiz-Cuaran, S., Swalduz, A., Foy, J. P., Marteau, S., Morel, A. P., Fauvet, F., ... & Saintigny, P. (2022). Epithelial-to-mesenchymal transition promotes immune escape by inducing CD70 in non-small cell lung cancer. European Journal of Cancer, 169, 106-122. https://doi.org/10.1016/j.ejca.2022.03.038
Pizzoni, A., Zhang, X., Naim, N., & Altschuler, D. L. (2023). Soluble cyclase-mediated nuclear cAMP synthesis is sufficient for cell proliferation. Proceedings of the National Academy of Sciences, 120(4), e2208749120. https://doi.org/10.1073/pnas.2208749120
Saikia, B. B., Bhowmick, S., Malat, A., Rani, M. P., Thaha, A., & Abdul-Muneer, P. M. (2024). ICAM-1 deletion using CRISPR/Cas9 protects the brain from traumatic brain injury-induced inflammatory leukocyte adhesion and transmigration cascades by attenuating the Paxillin/FAK-Dependent Rho GTPase Pathway. Journal of Neuroscience, 44(11). https://doi.org/10.1523/JNEUROSCI.1742-23.2024
Santo-Domingo, J., Lassueur, S., Galindo, A. N., Alvarez-Illera, P., Romero-Sanz, S., Caldero-Escudero, E., ... & Wiederkehr, A. (2023). SLC25A46 promotes mitochondrial fission and mediates resistance to lipotoxic stress in INS-1E insulin-secreting cells. Journal of Cell Science, 136(8), jcs260049. https://doi.org/10.1242/jcs.260049
Smeir, M., Chumala, P., Katselis, G. S., & Liu, L. (2024). Lymphocyte-Specific protein 1 regulates expression and stability of endothelial nitric oxide synthase. Biomolecules, 14(1), 111. https://doi.org/10.3390/biom14010111
Zhang, X., Tanwar, V. S., Jose, C. C., Lee, H. W., & Cuddapah, S. (2022). Transcriptional repression of E‐cadherin in nickel‐exposed lung epithelial cells mediated by loss of Sp1 binding at the promoter. Molecular carcinogenesis, 61(1), 99-110. https://doi.org/10.1002/mc.23364
This product has no review yet.
Controls and Related Product:
UNIT
40µg (250pmol/ 25µL)
UNIT
32.5µg (250pmol/ 25µL)
{"id":632,"cat_no":"G932","sku_in_m2":"","old_cat_no":"G932","name":"CRISPR Genomic Cleavage Detection Kit","category_id":208,"category_name":"Cas Proteins \u0026 CRISPR Screening","frontend_category_id":71,"frontend_category_ids":"86,57,67,71","gene_id":0,"gene_name":"","gene_full_name":"","alias":"","unit_quantity":"100 Reactions","accession_number":"","species_id":0,"species":"","raised_in_id":0,"raised_in":"","cell_type_id":0,"cell_type":"","tissue_id":0,"tissue":"","price":"170.0000","price_before_discount":"0.0000","keywords":"0","invisible_search_keywords":"crispr detection kit, gene editing detection kit, mutation detection kit, detection kit,","filter_group_id":34,"filter_group_name":"Molecular Biology","filter_id":41,"filter_name":"Kits \u0026 Other Enzymes","search_filter_ids":"32,34","search_filter_keys":"32-34","temp_field":null,"customization":0,"filter_table_id":18085025,"filter_table_name":"","related_group_id":17,"url":"","operator":"","points_rule":7,"status":1,"need_remove":0,"document_sort":"","price_old":"165.0000","price2024":"165.0000","created_at":null,"updated_at":"2025-03-05 23:25:06","Confirmed":"Tony Confirmed","geo_price_before_discount":"0.00","geo_titer7_price_before_discount":"149.00","cate_url":"cas9-proteins.html","frontendBreadcrumbs":[{"id":57,"pid":86,"name":"Genetic Materials","page_id":8,"page_type":"App\\Models\\Category","href":null,"sort":57,"show_in_nav":1,"show_in_homepage":1,"has_children":1,"status":1,"related_table":null,"created_at":null,"updated_at":"2025-01-20 08:46:24","url_key":"genetic-materials.html","url":"genetic-materials.html"},{"id":67,"pid":57,"name":"CRISPR","page_id":15,"page_type":"App\\Models\\Category","href":null,"sort":67,"show_in_nav":1,"show_in_homepage":1,"has_children":1,"status":1,"related_table":null,"created_at":null,"updated_at":null,"url_key":"CRISPR-Cas9-sgRNA.html","url":"CRISPR-Cas9-sgRNA.html"},{"id":71,"pid":67,"name":"Cas Proteins \u0026 CRISPR Screening","page_id":223,"page_type":"App\\Models\\Cms","href":null,"sort":71,"show_in_nav":1,"show_in_homepage":1,"has_children":0,"status":1,"related_table":null,"created_at":null,"updated_at":null,"url_key":"cas9-proteins.html","url":"cas9-proteins.html"}],"price_flag":0,"builderURL":"","builderViewerURL":"","mapURL":"","viewerURL":"","builderType":"","productType":"App\\Models\\CatalogBaseMolecular","geo_price":"170.00","geo_price_symbol":"$170.00","category":{"id":208,"categories":"Cas Proteins \u0026 CRISPR Screening","pid":15,"url_key":"cas9-proteins.html","css":null,"wid":0,"breadcrumbs":null,"breadcrumbsIdJson":"","breadcrumbsHtml":"","description":"\u003Cp\u003ECas9 proteins\u003C\/p\u003E\n\u003Cdiv id=\u0022gtx-trans\u0022 style=\u0022position: absolute; left: -35px; top: -10.5px;\u0022\u003E\n\u003Cdiv class=\u0022gtx-trans-icon\u0022\u003E\u003C\/div\u003E\n\u003C\/div\u003E","meta_title":"Cas Proteins \u0026 CRISPR Screening","meta_keywords":"Cas Proteins \u0026 CRISPR Screening","meta_description":"Cas Proteins \u0026 CRISPR Screening","deleted_at":null,"enable":"N","parent_list":"8,15","table_name":null,"image":null,"independentPage":0,"top_type":1,"sort_order":null,"in_footer":1,"fid":null,"created_at":"2024-08-28 23:24:53","updated_at":"2024-12-19 17:58:36"},"info":{"id":16605,"cat_no_base":"G932","parent_id":632,"description":"\u003Cp\u003E\u003Cstrong\u003Eabm\u003C\/strong\u003E\u0026rsquo;s \u003Cstrong\u003ECRISPR Genomic Cleavage Detection Kit\u003C\/strong\u003E is a \u003Cstrong\u003Esimple and rapid assay designed to verify your genomic editing process\u003C\/strong\u003E. Precision is essential in genome editing and gene modification, and confirming successful gene editing early can save both time and resources. This kit allows you to easily detect genomic cleavage, ensuring accurate results in your experiments.\u003C\/p\u003E\n\u003Cp\u003EThe kit uses \u003Cstrong\u003ECRISPR-edited samples as templates\u003C\/strong\u003E in PCR reactions targeting your specific region of interest. It requires a pair of primers flanking each sgRNA target site to detect genomic cleavage, which can be ordered from \u003Cstrong\u003Eabm\u003C\/strong\u003E (\u003Ca href=\u0022\/Primer-Design-Primer-Synthesis.html\u0022\u003ECat. No. C336\u003C\/a\u003E - \u003Cstrong\u003EsgRNA PCR Primer Pair Design \u0026 Synthesis Service)\u003C\/strong\u003E. After PCR amplification, the products are denatured and re-annealed, creating mismatches within the double-strand DNA. Our detection enzyme recognizes these mismatches and cleaves the strands to produce distinguishable band sizes upon gel analysis.\u003C\/p\u003E\n\u003Cp\u003EThe \u003Cstrong\u003Eready-to-use kit\u003C\/strong\u003E includes all necessary reagents, including a \u003Cstrong\u003Econtrol template\u003C\/strong\u003E and \u003Cstrong\u003Eprimers\u003C\/strong\u003E to ensure reliable results. The control template produces bands of approximately 750bp after PCR amplification and 500bp and 250bp after enzymatic cleavage. With a quick processing time, this assay is a valuable addition to any genomic-editing workflow.\u003C\/p\u003E\n\u003Cp\u003E\u003Cstrong\u003EKit Features:\u003C\/strong\u003E\u003C\/p\u003E\n\u003Cul\u003E\n\u003Cli\u003EEase of use with simple steps\u003C\/li\u003E\n\u003Cli\u003ERapid set-up\u003C\/li\u003E\n\u003Cli\u003EStreamlined protocol suitable for high-throughput applications\u003C\/li\u003E\n\u003C\/ul\u003E\n\u003Ctable class=\u0022bootstrap-table bootstrap-table-hover abm-service-table abm-perfect-table2\u0022 style=\u0022margin: 20px 0;\u0022\u003E\n\u003Cthead\u003E\n\u003Ctr\u003E\n\u003Cth\u003E\u003Cstrong\u003EProduct Component\u003C\/strong\u003E\u003C\/th\u003E\n\u003Cth\u003E\u003Cstrong\u003EQuantity\u003C\/strong\u003E\u003C\/th\u003E\n\u003C\/tr\u003E\n\u003C\/thead\u003E\n\u003Ctfoot\u003E\n\u003Ctr\u003E\n\u003Ctd\u003ECell Lysis Buffer\u003C\/td\u003E\n\u003Ctd\u003E1.25 ml\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EProtein Degarder\u003C\/td\u003E\n\u003Ctd\u003E100\u0026micro;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EDetection Enzyme\u003C\/td\u003E\n\u003Ctd\u003E50 \u0026mu;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003E10X Detection Buffer\u003C\/td\u003E\n\u003Ctd\u003E200 \u0026mu;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EControl Primer \u0026 Template\u003C\/td\u003E\n\u003Ctd\u003E10 \u0026mu;l\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003Ctr\u003E\n\u003Ctd\u003EMegaFi\u0026trade; Pro Fidelity 2X MasterMix\u003C\/td\u003E\n\u003Ctd\u003E2 x 1.25 ml\u003C\/td\u003E\n\u003C\/tr\u003E\n\u003C\/tfoot\u003E\n\u003C\/table\u003E","disclaimer":null,"application":null,"components":null,"cas9_origin":null,"concentration":null,"enzymes_size":null,"genecraft_series":null,"guarantee":null,"population":null,"qc":null,"format_general":null,"including_screening_kit":null,"expression_system_general":null,"purity":null,"image":"no_selection","insert_size":null,"shipping_conditions":null,"source_catalog_number":null,"inactivation_protocol":null,"led_viewer_compatibility":null,"unit_definition":null,"vector":null,"reaction_buffer":null,"storage_buffer":null,"caution":null,"storage_condition":"\u003Cp\u003EStore all components at -20\u00b0C.\u003C\/p\u003E","product_volume":null,"reporter":null,"safeview_series":null,"source_catno":null,"stain_color":null,"supplier":null,"internal_supplier":null,"internal_note":null,"inventory_location":"PCR-2A","note":null,"recommend":null,"depositor":null,"licensor_name":null,"licensor_contact_information":null,"contract_termination_date":null,"royalty_rates":null,"cas_type":null,"cas_origin":null,"cas_protein_marker":null,"source":null,"endotoxin_level":null,"additional_information":null,"titer":null,"nucleotide_format":null,"protocol_overview":null,"source_price":null,"created_at":"2022-09-09 06:29:47","updated_at":"2025-03-05 23:25:06","short_description":"\u003Cp\u003EThe CRISPR Genomic Cleavage Detection Kit is designed as a simple, rapid assay to verify\u00a0your genomic editing process.\u003C\/p\u003E","reaction_definition":null,"specificity":null,"promotions":null},"maps":[],"media":[{"id":379752,"parent_id":632,"parent_type":"App\\Models\\CatalogBaseMolecular","file_path":"\/c\/r\/crispr-genomic-cleavage-detection-kit-g932.jpg","title":null,"text":null,"file_type":"image","alt":null,"url":null,"position":1,"status":0,"entity_id_m2":18085025,"sku_in_m2":"G932","value_id_m2":49912342,"attribute_id":90,"created_at":"2022-07-19 04:36:22","updated_at":"2024-03-26 20:42:00"},{"id":379751,"parent_id":632,"parent_type":"App\\Models\\CatalogBaseMolecular","file_path":"\/g\/9\/g932-crispr-genomic-cleavage-detection-kit_workflow.jpg","title":null,"text":null,"file_type":"image","alt":null,"url":null,"position":2,"status":0,"entity_id_m2":18085025,"sku_in_m2":"G932","value_id_m2":49912336,"attribute_id":90,"created_at":"2022-07-19 04:36:22","updated_at":"2025-03-05 23:25:37"},{"id":384867,"parent_id":632,"parent_type":"App\\Models\\CatalogBaseMolecular","file_path":"\/upload\/xHc4bbfjtBO1UBRGiJRAytZbh0KIoJG67pPpc3sH.png","title":null,"text":null,"file_type":"image","alt":null,"url":null,"position":3,"status":1,"entity_id_m2":null,"sku_in_m2":null,"value_id_m2":null,"attribute_id":0,"created_at":"2024-03-26 20:41:35","updated_at":"2025-03-05 23:25:37"},{"id":379750,"parent_id":632,"parent_type":"App\\Models\\CatalogBaseMolecular","file_path":"\/g\/9\/g932-crispr-genomic-cleavage-detection-kit_data1.jpg","title":null,"text":null,"file_type":"image","alt":null,"url":null,"position":4,"status":0,"entity_id_m2":18085025,"sku_in_m2":"G932","value_id_m2":49912335,"attribute_id":90,"created_at":"2022-07-19 04:36:22","updated_at":"2024-03-26 20:42:00"}],"gene":null}