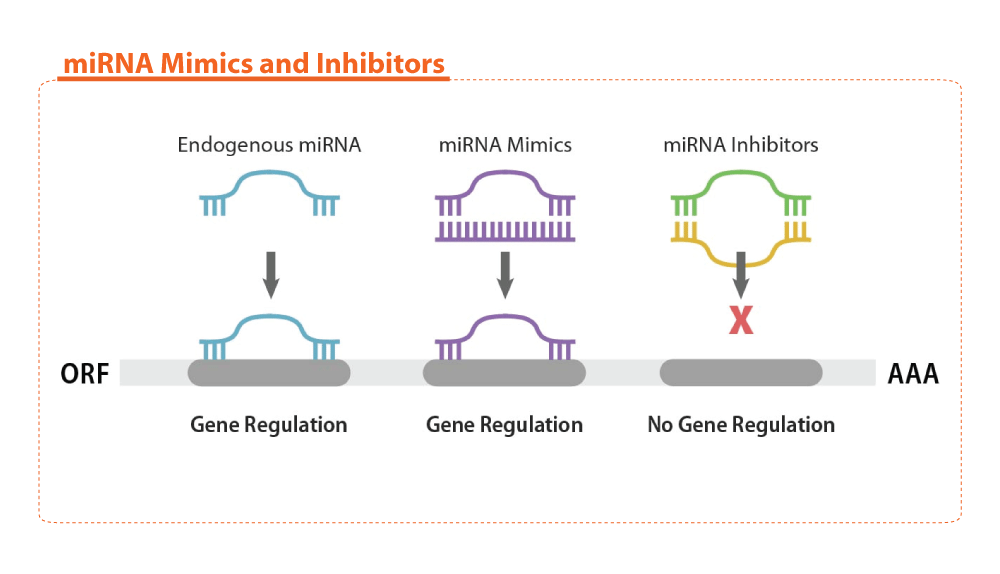
Synthetic miRNA Inhibitors and Antagomirs
abm offers miRNA Inhibitors and Antagomirs that can be used for miRNA loss-of-function studies, target validation, and screening experiments. miRNA Inhibitors are complementary antisense single-stranded oligos which can bind to endogenous mature miRNA, while Antagomirs are a type of miRNA Inhibitor with chemical modifications to prevent degradation and increase transfection efficiency. Both Inhibitors and Antagomirs block miRNA regulation of target gene expression by stable suppression and can knock down native miRNA expression in cells.
Key Features:
- We offer a collection of Inhibitors & Antagomirs targeting >6000 miRNAs from human, mouse, and rat.
- Synthetic miRNA inhibitors are easily transfected into many cell types for rapid knockdown of mature miRNA.
- Antagomirs have chemical modifications to increase transfection efficiency and stability.
Search our collection of miRNA Technology products using miRNA name, symbol or accession number:
Search results will be displayed here
Additional Information
Documents
Top Publications
| 01 | MicroRNA-377 Regulates Mesenchymal Stem Cell-Induced Angiogenesis in Ischemic Hearts by Targeting VEGF. Wen, Z et al. Plos One 9 (9):e104666 (2014). doi:10.1371/journal.pone.0104666 |
| 02 | MicroRNA-146a induces immune suppression and drug-resistant colorectal cancer cells. Tumor Biology 39(5):1-12 (2017). doi:10.1177/1010428317698365 |
| 03 | Down-Regulation of eIF4GII by miR-520c-3p Represses Diffuse Large B Cell Lymphoma Development. Mazan-Mamczarz, K et al. PLoS Genet. 10(1):e1004105. (2014) doi: 10.1371/journal.pgen.1004105 |
FAQs
| What is the Sanger miRBase Sequence Database? |
|
miRBase is a sequence database that has been established by the Sanger Institute. Each entry in the microRNA Registry represents a predicted hairpin portion of a microRNA transcript (termed mir in the database), with information on the location and sequence of the mature microRNA sequence (termed miR). The database provides microRNA gene hunters with unique names for novel microRNA genes prior to publication of results and a searchable database of published microRNA.
|
| When was the latest update of your array sequences? |
|
The content of our arrays has been updated to miRBase Release 16.0.
|
| Which genes are targeted by a specific miRNA? |
|
You can search for predicted and validated miRNA target genes at http://mirbase.org/.
Just type in your miRNA name (eg. hsa-mir-145) or accession number in the search bar and look for the targeted genes. |
| Do I need to use an internal control primer for quantitative detection of miRNAs (the same as housekeeping primers which we use in qPCR for mRNAs detection)? |
|
For human miRNA plates, the controls are as follows:
1) SNORD44 Primers 2) SNORD48 Primers 3) U6 Primers 4) hsa-miR-16-5p Primers |