All-In-One 5X RT MasterMix with gDNA Removal
| Cat. No. | G592 | ||||||||
| Name | All-In-One 5X RT MasterMix with gDNA Removal | ||||||||
| Unit | 100 rxn | ||||||||
| Category | Reverse Transcriptase & RT-PCR | ||||||||
| Description |
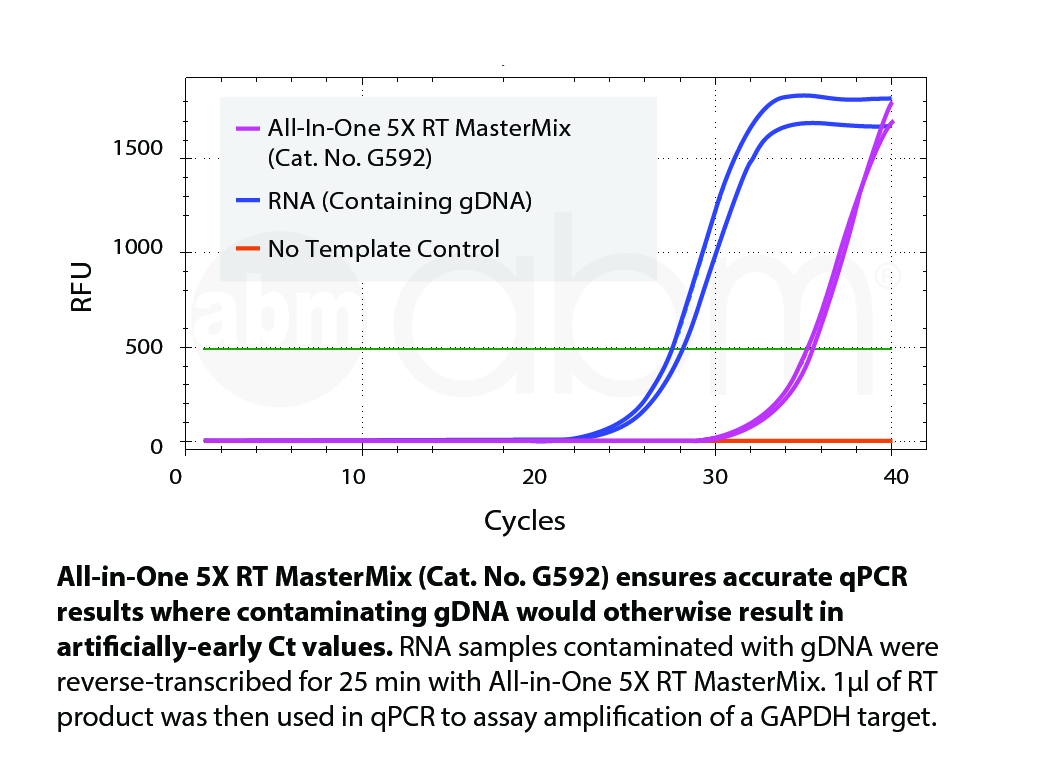
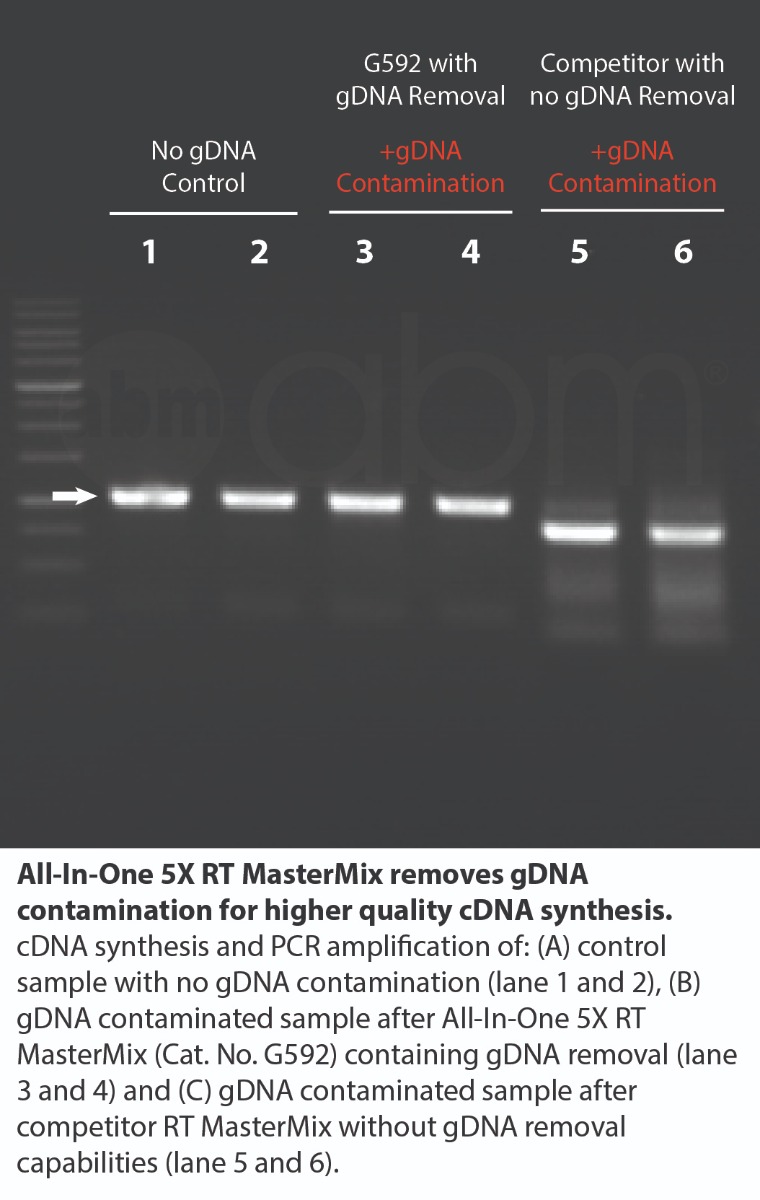
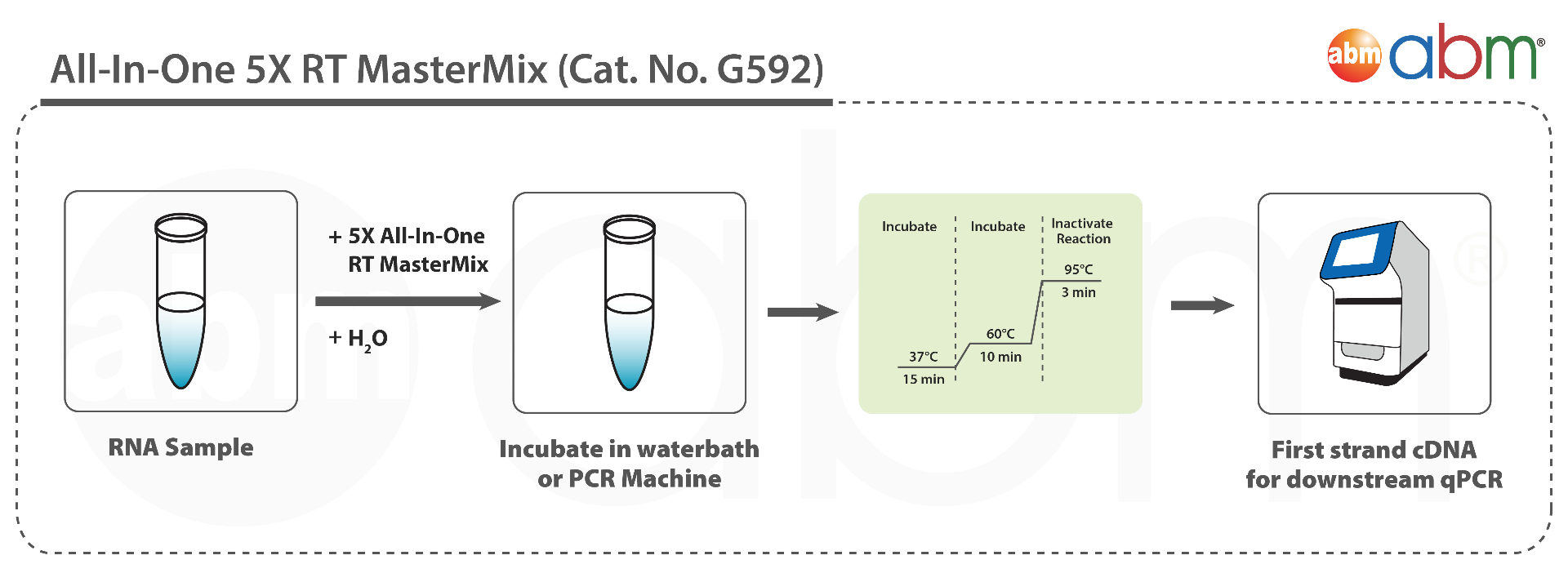
The All-In-One 5X RT MasterMix with gDNA Removal offers a convenient, ready-to-use solution for first-strand cDNA synthesis with genomic DNA removal, all in a single tube. This powerful formulation addresses the common challenge of genomic DNA contamination, ensuring accurate RNA detection without compromising reverse transcription or cDNA synthesis. The optimized MasterMix includes abm’s proprietary OneScript® Hot Reverse Transcriptase, RNaseOFF Ribonuclease Inhibitor, temperature-sensitive DNase, dNTPs, and a carefully balanced mix of Oligo (dT)s and Random Primers. By eliminating the need for separate reagents, it provides a straightforward, reliable, and reproducible cDNA synthesis process with high yields—even from difficult samples. The high-quality cDNA produced is suitable for a wide range of downstream applications. Product Features:
|
||||||||
| Application |
|
||||||||
| Concentration | 5X | ||||||||
| Storage Condition |
Store at -20°C. |
||||||||
| Note |
For applications using pure RNA where genomic DNA removal is not required, try our OneScript® Hot 5X RT MasterMix (Cat. No. G590). It offers a faster protocol for an even more efficient cDNA synthesis. |
||||||||
| Material Citation | If use of this material results in a scientific publication, please cite the material in the following manner: Applied Biological Materials Inc, Cat. No. G592 |
| What's included in this kit? | |
|
The kit includes everything you need for cDNA synthesis in one tube: All-In-One 5X RT MasterMix (100 reactions, 400 µl), which contains reverse transcriptase, RNaseOFF RNase inhibitor, DNase, dNTPs, and primers. Additionally, a tube of Nuclease-Free H₂O (1.0 ml) is provided to complete your reaction setup.
|
| How should I store the All-In-One 5X RT MasterMix with gDNA Removal? | |
|
Store the All-In-One 5X RT MasterMix with gDNA Removal at -20°C. This product is unstable if stored improperly. Reduce freeze-thaw cycling and handle with care.
|
| Can I use both total RNA and poly(A)+ mRNA for cDNA synthesis? | |
|
Yes, both total RNA and poly(A)+ mRNA can be used, though poly(A)+ mRNA typically yields higher quantities and better purity.
|
| How can I enhance genomic DNA removal? | |
|
To improve genomic DNA removal, increase the 37°C incubation time from 15 minutes to 30 minutes.
|
| Can I synthesize cDNA from long RNA transcripts? | |
|
Yes, for longer RNA transcripts, extend the 60°C incubation time to up to 30 minutes.
|
| What if genomic DNA removal isn’t necessary? | |
|
If genomic DNA removal is not required, use the OneScript® Hot 5X RT MasterMix (Cat. No. G590) for a faster and more efficient cDNA synthesis process.
|
| What can I use the synthesized cDNA for? | |
|
The high-quality cDNA can be used in a variety of downstream applications, including gene expression analysis, cloning, and PCR-based assays. BlasTaq™ 2X qPCR Master Mix (Cat. No. G891) is well suited to downstream qPCR applications.
|
| How should I store the synthesized cDNA? | |
|
Store the synthesized first-strand cDNA at -20°C for long-term use.
|
| Why am I getting low cDNA yields? | |
|
Low cDNA yields can be caused by poor RNA integrity, contamination, or insufficient RNA input. To improve yield:
|
| How much RNA template should I use for cDNA synthesis? | |
|
We recommend using 1 ng to 2 μg of RNA per reaction.
|
| What volumes of cDNA should I use for downstream PCR? | |
|
Typically, use 1 μl of cDNA in a 25 μl PCR reaction. You can add up to 20% of the PCR volume (e.g., 5 μl in a 25 μl PCR), depending on your target and primers.
|
|
Do I need an RNase inhibitor when using this product?
|
|
|
All of abm's Reverse Transcriptases include RNaseOFF Ribonuclease Inhibitor, which protects RNA from degradation and is resistant to oxidation, even under low DTT concentrations. Therefore, there’s no need to add an external RNase inhibitor, as RNaseOFF provides optimal RNA protection during reverse transcription.
|
| What discontinued abm products is Cat. No. G592 equivalent to? | |
|
This product is abm's next generation of RT MasterMixes and is functionally equivalent to Cat. No. G485, G486, G490, G492, G916 (5X All-In-One RT MasterMixes with AccuRT Genomic DNA Removal Kit or with ExCellenCT Lysis Kits), and G451, G452, G453, G454 (OneScript® cDNA Synthesis SuperMixes), with improved performance. For more information, contact our customer service team at technical@abmgood.com.
|
-
Accogli, A., Lu, S., Musante, I., Scudieri, P., Rosenfeld, J. A., Severino, M., ... & Salpietro, V. (2023). Loss of neuron navigator 2 impairs brain and cerebellar development. The Cerebellum, 22(2), 206-222. https://doi.org/10.1007/s12311-022-01379-3
An, H., Yan, C., Ma, J., Gong, J., Gao, F., Ning, C., ... & Yu, Q. (2023). Immune inhibitory receptor-mediated immune response, metabolic adaptation, and clinical characterization in patients with COVID-19. Scientific Reports, 13(1), 19221. https://doi.org/10.1038/s41598-023-45883-wAn, L., Zhai, Q., Tao, K., Xiong, Y., Ou, W., Yu, Z., ... & Lu, M. (2024). Quercetin induces itaconic acid-mediated M1/M2 alveolar macrophages polarization in respiratory syncytial virus infection. Phytomedicine, 130, 155761. https://doi.org/10.1016/j.phymed.2024.155761Baker, K. M., Hoda, S., Saha, D., Georgescu, L., Serratore, N. D., Zhang, Y., ... & Briggs, S. D. (2021). Set1-mediated histone H3K4 methylation is required for azole induction of the ergosterol biosynthesis genes and antifungal drug resistance in Candida glabrata. bioRxiv, 2021-11. https://doi.org/10.1101/2021.11.17.469015Banfi, C., Brioschi, M., Vicentini, L. M., & Cattaneo, M. G. (2022). The Effects of Silencing PTX3 on the Proteome of Human Endothelial Cells. International Journal of Molecular Sciences, 23(21), 13487. https://doi.org/10.3390/ijms232113487Basile, A., Zannella, C., De Marco, M., Sanna, G., Franci, G., Galdiero, M., ... & Marzullo, L. (2023). Spike-mediated viral membrane fusion is inhibited by a specific anti-IFITM2 monoclonal antibody. Antiviral Research, 211, 105546. https://doi.org/10.1016/j.antiviral.2023.105546Besse, S., Sakaguchi, T., Gauthier, L., Sahaf, Z., Péloquin, O., Gonzalez, L., ... & Serohijos, A. W. (2023). Genetic landscape of an in vivo protein interactome. bioRxiv, 2023-12. https://doi.org/10.1101/2023.12.14.571726Bi, J., Fu, X., Jiang, Y., Wang, J., Li, D., Xiao, M., & Mou, H. (2025). Low molecular weight galactomannan alleviates diarrhea induced by senna leaf in mice via intestinal barrier improvement and gut microbiota modulation. Food & Function. https://doi.org/10.1039/D4FO04375HBibbò, F., Asadzadeh, F., Boccia, A., Sorice, C., Bianco, O., Saccà, C. D., ... & Zollo, M. (2024). T argeting group 3 medulloblastoma by the anti-PRUNE-1 and anti-LSD1/KDM1A epigenetic molecules. International Journal of Molecular Sciences, 25(7), 3917. https://doi.org/10.3390/ijms25073917Bolf, E. L., Gillis, N. E., Barnum, M. S., Beaudet, C. M., Yu, G. Y., Tomczak, J. A., ... & Carr, F. E. (2020). The thyroid hormone receptor-RUNX2 Axis: a novel tumor suppressive pathway in breast cancer. Hormones and Cancer, 11, 34-41. https://doi.org/10.1007/s12672-019-00373-2Bolf, E. L., Gillis, N. E., Davidson, C. D., Rodriguez, P. D., Cozzens, L., Tomczak, J. A., ... & Carr, F. E. (2020). Thyroid hormone receptor beta induces a tumor-suppressive program in anaplastic thyroid cancer. Molecular Cancer Research, 18(10), 1443-1452. https://doi.org/10.1158/1541-7786.MCR-20-0282Camacho-Jiménez, L., Leyva-Carrillo, L., Gómez-Jiménez, S., & Yepiz-Plascencia, G. (2024). Naphthalene and phenanthrene affect differentially two glutathione S-transferases (GSTs) expression, GST activity, and glutathione content in white shrimp P. vannamei. Aquatic Toxicology, 107005. https://doi.org/10.1016/j.aquatox.2024.107005Castiglioni, S., Pezzoli, L., Pezzani, L., Lettieri, A., Di Fede, E., Cereda, A., ... & Gervasini, C. (2024). Expanding the clinical spectrum of PPP3CA variants-alternative isoforms matter. Orphanet Journal of Rare Diseases, 19(1), 481. https://doi.org/10.1186/s13023-024-03507-0Cen, C., Chen, J., Lin, L., Chen, M., Dong, F., Shen, Z., ... & Gao, F. (2022). Fancb deficiency causes premature ovarian insufficiency in mice. Biology of Reproduction, 107(3), 790-799. https://doi.org/10.1093/biolre/ioac103Chen, J., Lin, X., Xiang, W., Chen, Y., Zhao, Y., Huang, L., & Liu, L. (2024). DNA target binding-induced pre-crRNA processing in type II and V CRISPR-Cas systems. Nucleic Acids Research, gkae1241. https://doi.org/10.1093/nar/gkae1241Chen, K., Wang, Y., Wu, C., Du, Y., Tang, H., Zheng, S., ... & Wu, G. (2024). ln vitro assessment of immunomodulatory and osteogenic properties in 3D-printed hydroxyapatite/barium titanate piezoelectric ceramic scaffolds. Ceramics International, 50(6), 8751-8759. https://doi.org/10.1016/j.ceramint.2023.12.192Chen, L., Qin, Z., & Ruan, Z. B. (2023). Hyperoside alleviates doxorubicin-induced myocardial cells apoptosis by inhibiting the apoptosis signal-regulating kinase 1/p38 pathway. PeerJ, 11, e15315. https://doi.org/10.7717/peerj.15315Chen, M., Dong, F., Shen, Z., Wu, H., Cen, C., Cui, X., ... & Gao, F. (2021). PRMT5 regulates ovarian follicle development by facilitating Wt1 translation. Elife, 10, e68930. https://doi.org/10.7554/eLife.68930Chen, R., Yang, M., Tu, Z., Xie, F., Chen, J., Luo, T., ... & He, C. (2022). Eukaryotic translation initiation factor 4E family member nCBP facilitates the accumulation of TGB-encoding viruses by recognizing the viral coat protein in potato and tobacco. Frontiers in Plant Science, 13, 946873. https://doi.org/10.3389/fpls.2022.946873Chen, T., Xiang, L., Zhang, W., Xia, Z., & Chen, W. (2024). AGXT2 Suppresses the Proliferation and Dissemination of Hepatocellular Carcinoma Cells by Modulating Intracellular Lipid Metabolism. Journal of Hepatocellular Carcinoma, 1623-1639. https://doi.org/10.2147/JHC.S470250Chen, X. C., Huang, L. F., Tang, J. X., Wu, D., An, N., Ye, Z. N., ... & Yang, C. (2023). Asiatic acid alleviates cisplatin-induced renal fibrosis in tumor-bearing mice by improving the TFEB-mediated autophagy-lysosome pathway. Biomedicine & Pharmacotherapy, 165, 115122. https://doi.org/10.1016/j.biopha.2023.115122Chen, X., Liu, Y., Luo, X., Pan, T., Zhang, T., Hu, L., ... & Wei, F. (2024). HPV16 E6-induced M2 macrophage polarization in the cervical microenvironment via exosomal miR-204-5p. Scientific Reports, 14(1), 23725. https://doi.org/10.1038/s41598-024-74399-0Cheng, D., Zhou, D., Wang, Y., Wang, B., He, Q., Song, B., & Chen, H. (2021). Ralstonia solanacearum type III effector RipV2 encoding a novel E3 ubiquitin ligase (NEL) is required for full virulence by suppressing plant PAMP-triggered immunity. Biochemical and Biophysical Research Communications, 550, 120-126. https://doi.org/10.1016/j.bbrc.2021.02.082Chung, H. L., Rump, P., Lu, D., Glassford, M. R., Mok, J. W., Fatih, J., ... & Harel, T. (2022). De novo variants in EMC1 lead to neurodevelopmental delay and cerebellar degeneration and affect glial function in Drosophila. Human Molecular Genetics, 31(19), 3231-3244. https://doi.org/10.1093/hmg/ddac053Claudia, M. V., Javiera, A. A., Sebastián, N. S., José, F. R., & Gloria, L. (2024). Interplay between desiccation and oxidative stress responses in iron-oxidizing acidophilic bacteria. Journal of Biotechnology, 383, 64-72. https://doi.org/10.1016/j.jbiotec.2024.01.017Cocci, P., Angeletti, M., Mosconi, G., Olivotto, I., Zarantoniello, M., & Palermo, F. A. (2024). Replacement of fish meal with full fat Hermetia illucens modulates hepatic FXR signaling in juvenile rainbow trout (Oncorhynchus mykiss): Exploring a potential role of ecdysteroids. Heliyon, 10(22). https://doi.org/10.1016/j.heliyon.2024.e40302Cocci, P., Mazzocchi, V., Marconi, M., Mosconi, G., & Palermo, F. A. (2024). Assessing the impact of weathered polystyrene collected from the marine environment on oxidative stress responses in Zophobas morio larvae: A preliminary study. Environmental Advances, 17, 100593. https://doi.org/10.1016/j.envadv.2024.100593Davidson, C. D., Bolf, E. L., Gillis, N. E., Cozzens, L. M., Tomczak, J. A., & Carr, F. E. (2021). Thyroid hormone receptor beta inhibits PI3K-Akt-mTOR signaling axis in anaplastic thyroid cancer via genomic mechanisms. Journal of the Endocrine Society, 5(8), bvab102. https://doi.org/10.1210/jendso/bvab102de Antonellis, P., Ferrucci, V., Miceli, M., Bibbo, F., Asadzadeh, F., Gorini, F., ... & Zollo, M. (2024). Targeting ATP2B1 impairs PI3K/Akt/FOXO signaling and reduces SARS-COV-2 infection and replication. EMBO reports, 1-34. https://doi.org/10.1038/s44319-024-00164-zDi Fede, E., Lettieri, A., Taci, E., Castiglioni, S., Rebellato, S., Parodi, C., ... & Gervasini, C. (2024). Characterization of a novel HDAC2 pathogenetic variant: a missing puzzle piece for chromatinopathies. Human Genetics, 1-13. https://doi.org/10.1007/s00439-024-02675-0Dong, H., Yang, L., Liu, Y., Tian, G., Tang, H., Xin, S., ... & Qian, W. (2023). Detection of new candidate genes controlling seed weight by integrating gene coexpression analysis and QTL mapping in Brassica napus L. The Crop Journal, 11(3), 842-851. https://doi.org/10.1016/j.cj.2022.09.009Dong, K., Geng, C., Zhan, X., Sun, Z., Pu, Q., Li, P., ... & Gao, H. (2023). GREB1L overexpression is associated with good clinical outcomes in breast cancer. European Journal of Medical Research, 28(1), 510. https://doi.org/10.1186/s40001-023-01483-yDorajoo, R., Ihsan, M. O., Liu, W., Lim, H. Y., Angeli, V., Park, S. J., ... & Sorokin, V. A. (2023). Vascular smooth muscle cells in low SYNTAX scores coronary artery disease exhibit proinflammatory transcripts and proteins correlated with IL1B activation. Atherosclerosis, 365, 15-24. https://doi.org/10.1016/j.atherosclerosis.2022.12.005Dort, J., Orfi, Z., Fiscaletti, M., Campeau, P. M., & Dumont, N. A. (2023). Gpr18 agonist dampens inflammation, enhances myogenesis, and restores muscle function in models of Duchenne muscular dystrophy. Frontiers in Cell and Developmental Biology, 11, 1187253. https://doi.org/10.3389/fcell.2023.1187253Elbialy, A., Kitauchi, M., & Yamanouchi, D. (2023). Antioxidants and azd0156 Rescue Inflammatory Response in Autophagy-Impaired Macrophages. International journal of molecular sciences, 25(1), 169. https://doi.org/10.3390/ijms25010169Eyal, Y., & Carmi, N. (2025). A 2OGD multi-gene cluster encompasses functional and tissue specificity that direct furanocoumarin and pyranocoumarin biosynthesis in citrus. New Phytologist, 245, 1547-1562. https://doi.org/10.1111/nph.20322Fong-McMaster, C., Pulente, S. M., Kennedy, L., Smith, T. K., Myers, S., Kanaan, M., ... & Harper, M. E. (2024). OPA1 mediates cardiac function and metabolism: in silico and in vivo evidence. bioRxiv, 2024-08. https://doi.org/10.1101/2024.08.23.605375Gallard, C., Lebsir, N., Khursheed, H., Reungoat, E., Plissonnier, M. L., Bré, J., ... & Grigorov, B. (2022). Heparanase-1 is upregulated by hepatitis C virus and favors its replication. Journal of hepatology, 77(1), 29-41. https://doi.org/10.1016/j.jhep.2022.01.008Gao, Y., Gong, Y., Lu, J., Yang, Y., Zhang, Y., Xiong, Y., & Shi, X. (2024). Dihydroartemisinin breaks the positive feedback loop of YAP1 and GLUT1-mediated aerobic glycolysis to boost the CD8+ effector T cells in hepatocellular carcinoma. Biochemical Pharmacology, 225, 116294. https://doi.org/10.1016/j.bcp.2024.116294Gonzalez Lopez, L. (2023). Consequences of local and global chromatin mechanics to adaption and genome stability in the budding yeast Saccharomyces cerevisiae. https://hdl.handle.net/1866/28568González, L., Kolbin, D., Trahan, C., Jeronimo, C., Robert, F., Oeffinger, M., ... & Michnick, S. W. (2023). Adaptive partitioning of a gene locus to the nuclear envelope in Saccharomyces cerevisiae is driven by polymer-polymer phase separation. Nature Communications, 14(1), 1135. https://doi.org/10.1038/s41467-023-36391-6Gostyński, A., Diercks, G. F., Escamez, M. J., Chandran, N. S., de Lucas, R., Garcia-Martin, A., ... & Pasmooij, A. M. (2022). Natural occurrence of autoantibodies against basement membrane proteins in epidermolysis bullosa. Journal of Investigative Dermatology, 142(7), 2014-2019. https://doi.org/10.1016/j.jid.2021.10.030Gregor, J. B., Gutierrez-Schultz, V. A., Hoda, S., Baker, K. M., Saha, D., Burghaze, M. G., ... & Briggs, S. D. (2023). An expanded toolkit of drug resistance cassettes for Candida glabrata, Candida auris, and Candida albicans leads to new insights into the ergosterol pathway. Msphere, 8(6), e00311-23. https://doi.org/10.1128/msphere.00311-23Gregor, J. B., Gutierrez-Schultz, V. A., Hoda, S., Baker, K. M., Saha, D., Burghaze, M. G., & Briggs, S. D. (2023). Expanding the toolkit for genetic manipulation and discovery in Candida species using a CRISPR ribonucleoprotein-based approach. bioRxiv. https://doi.org/10.1101/2023.06.16.545382Guo, X., Zhang, M., Feng, Y., Liu, X., Wang, C., Zhang, Y., ... & Guo, Y. (2024). Transcriptome analysis of salivary glands of rabies-virus-infected mice. Frontiers in Microbiology, 15, 1354936. https://doi.org/10.3389/fmicb.2024.1354936Guo, Y., Qin, J., Sun, R., Hao, P., Jiang, Z., Wang, Y., ... & Zhang, W. (2024). Molecular hydrogen promotes retinal vascular regeneration and attenuates neovascularization and neuroglial dysfunction in oxygen-induced retinopathy mice. Biological Research, 57. https://doi.org/10.1186/s40659-024-00515-zGuo, Y., Wei, C., Ding, H., Li, P., Gao, Y., Zhong, K., ... & Hu, J. (2024). Effects of cold stress on the blood-brain barrier in Plectropomus leopardus. BMC genomics, 25(1), 1031. https://doi.org/10.1186/s12864-024-10943-6Han, H., Zhou, Z., Shang, T., Li, S., Shen, X., Fang, J., & Cui, L. (2024). Silk Fibroin-Laponite Porous Microspheres as Cell Microcarriers for Osteogenic Differentiation. Tissue Engineering Part A. https://doi.org/10.1089/ten.tea.2024.0070Han, M. M., He, X. Y., Tang, L., Qi, L., Yang, M. Y., Wang, Y., ... & Jiang, H. L. (2023). Nanoengineered mesenchymal stem cell therapy for pulmonary fibrosis in young and aged mice. Science advances, 9(29), eadg5358. https://doi.org/10.1126/sciadv.adg5358Han, M., Liu, Y., Jin, C., Wang, X., Song, W., & Zhang, Q. (2022). Genome-wide identification, characterization and expression profiling of TRAF family genes in Sebastes schlegelii. Fish & Shellfish Immunology, 127, 203-210. https://doi.org/10.1016/j.fsi.2022.06.021Han, Z., Migheli, Q., & Kong, Q. (2024). Fusion Expression of Peptides with AflR Binuclear Zinc Finger Motif and Their Enhanced Inhibition of Aspergillus flavus: A Study of Engineered Antimicrobial Peptides. Journal of Agricultural and Food Chemistry. https://doi.org/10.1021/acs.jafc.4c01259Hao, R., Li, H., Li, X., Liu, J., Ji, X., Zhang, H., ... & Zhai, Z. (2025). Transcriptomic profiling of lncRNAs and mRNAs in a venous thrombosis mouse model. iScience, 28(2). https://doi.org/10.1016/j.isci.2024.111561Hernández-Adasme, C., Silva, H., Peña, Á., Vargas-Martínez, M. G., Salazar-Parra, C., Sun, B., & Escalona Contreras, V. (2024). Modifying the Ambient Light Spectrum Using LED Lamps Alters the Phenolic Profile of Hydroponically Grown Greenhouse Lettuce Plants without Affecting Their Agronomic Characteristics. Plants, 13(17), 2466. https://doi.org/10.3390/plants13172466Hiltunen, H., Hanani, H., Luoto, R., Turjeman, S., Ziv, O., Isolauri, E., ... & Rautava, S. (2021). Preterm infant meconium microbiota transplant induces growth failure, inflammatory activation, and metabolic disturbances in germ-free mice. Cell Reports Medicine, 2(11). https://doi.org/10.1016/j.xcrm.2021.100447Holland, S. H., Carmona-Martinez, R., O’Connor, K., O’Neil, D., Roos, A., Spendiff, S., & Lochmüller, H. (2024). A Deficiency in Glutamine-Fructose-6-Phosphate Transaminase 1 (Gfpt1) in Skeletal Muscle Results in Reduced Glycosylation of the Delta Subunit of the Nicotinic Acetylcholine Receptor (AChRδ). Biomolecules, 14(10), 1252. https://doi.org/10.3390/biom14101252Hu, C., Li, H., Tong, C., Zhang, D., & Lu, Y. (2024). Integrated transcriptomic and metabolomic analyses reveal the effect of mycorrhizal colonization on trifoliate orange root hair. Scientia Horticulturae, 336, 113429. https://doi.org/10.1016/j.scienta.2024.113429Hu, J., Wang, B., Yang, T., Li, N., Yang, H., Yu, Q., & Wang, J. (2022). A calcium-dependent protein kinase gene SpCPK33 from Solanum pennellii associated with increased cold tolerance in tomato. Journal of Plant Physiology, 279, 153834. https://doi.org/10.1016/j.jplph.2022.153834Hu, M., Zhou, J., Qiu, L., Song, R., Qin, X., Tan, Z., ... & Wang, X. (2024). Effects of soy protein on alleviating iron deficiency anemia in suckling rats with different iron supplements. Food Bioscience, 61, 104555. https://doi.org/10.1016/j.fbio.2024.104555Hu, Y., Shi, H., Ma, X., Xia, T., Wu, Y., Chen, L., ... & Li, X. (2023). Highly stable fibronectin-mimetic-peptide-based supramolecular hydrogel to accelerate corneal wound healing. Acta biomaterialia, 159, 128-139. https://doi.org/10.1016/j.actbio.2023.01.047Huang, R., Bie, S., Li, S., Yuan, B., Zhang, L., Zhang, Z., ... & Zhang, D. (2024). Strigolactones Negatively Regulate Tobacco Mosaic Virus Resistance in Nicotiana benthamiana. International Journal of Molecular Sciences, 25(15), 8518. https://doi.org/10.3390/ijms25158518Huang, T., Fei, M., Zhou, X., He, K., Yang, S., & Zhao, A. (2024). Effects of Different Photoperiods on the Transcriptome of the Ovary and Small White Follicles in Zhedong White Geese. Animals, 14(18), 2747. https://doi.org/10.3390/ani14182747Ji, T., Fang, B., Jin, Y., Zheng, C., Yuan, X., Dong, J., ... & Wu, F. (2024). Euglena Attenuates High-Fat-Diet-Induced Obesity and Especially Glucose Intolerance. Nutrients, 16(21), 3780. https://doi.org/10.3390/nu16213780Ji, T., Fang, B., Wu, F., Liu, Y., Cheng, L., Li, Y., ... & Zhu, L. (2023). Diet change improves obesity and lipid deposition in high-fat diet-induced mice. Nutrients, 15(23), 4978. https://doi.org/10.3390/nu15234978Ji, T., Fang, B., Zhang, M., & Liu, Y. (2023). Succinate Enhances Lipolysis and Decreases Adipocytes Size in Both Subcutaneous and Visceral Adipose Tissue from High-Fat-Diet-Fed Obese Mice. Foods, 12(23), 4285. https://doi.org/10.3390/foods12234285Ji, T., Fang, B., Zhang, M., & Liu, Y. (2023). Supplement of Succinate Reduces Lipid Deposition and Improves Metabolic Function in Obese Mice. https://doi.org/10.20944/preprints202304.0208.v1Ji, Y., Sun, C., & Wu, S. (2024). Transcriptomic and Biochemical Analysis of the Antimicrobial Mechanism of Lipopeptide Iturin W against Staphylococcus aureus. International Journal of Molecular Sciences, 25(18), 9949. https://doi.org/10.3390/ijms25189949Ji, Z., Liu, Z., Han, Y., & Sun, Y. (2022). Exogenous dopamine promotes photosynthesis and carbohydrate metabolism of downy mildew-infected cucumber. Scientia Horticulturae, 295, 110842. https://doi.org/10.1016/j.scienta.2021.110842Jiang, K., Zhang, F., Chen, Y., Li, X., Zhao, X., Jiang, P., & Li, Y. (2024). Fosfenopril Attenuates Inflammatory Response in Diabetic Dry Eye Models by Inhibiting the TLR4/NF-κB/NLRP3 Signaling Pathway. Investigative Ophthalmology & Visual Science, 65(6), 2-2. https://doi.org/10.1167/iovs.65.6.2Jin, C., Yan, K., Wang, M., Song, W., Kong, X., & Zhang, Z. (2023). Identification, characterization and functional analysis of fibroblast growth factors in black rockfish (Sebastes schlegelii). International Journal of Molecular Sciences, 24(4), 3626. https://doi.org/10.3390/ijms24043626Kang, Y., Liang, S., Gao, S., & Chen, J. (2024). Generation of a hiPSC line (TONGJIi001-A) from a 46, XX, ins (1; 15)(p13. 3; q22. 31q26. 1), inv (2)(p22. 1p16. 3), t (2; 14)(q34; q12) infertility patient. Stem Cell Research, 81, 103561. https://doi.org/10.1016/j.scr.2024.103561Khan, F., Lin, Y., Ali, H., Pang, L., Dunterman, M., Hsu, W. H., ... & Chen, P. (2023). LDHA-regulated tumor-macrophage symbiosis promotes glioblastoma progression. Research Square. https://doi.org/10.21203/rs.3.rs-3401154/v1Khan, F., Lin, Y., Ali, H., Pang, L., Dunterman, M., Hsu, W. H., ... & Chen, P. (2024). Lactate dehydrogenase A regulates tumor-macrophage symbiosis to promote glioblastoma progression. Nature communications, 15(1), 1987. https://doi.org/10.1038/s41467-024-46193-zKopinski-Grünwald, O., Guillaume, O., Ferner, T., Schädl, B., & Ovsianikov, A. (2024). Scaffolded spheroids as building blocks for bottom-up cartilage tissue engineering show enhanced bioassembly dynamics. Acta Biomaterialia, 174, 163-176. https://doi.org/10.1016/j.actbio.2023.12.001Kremer, R., Mihalcioiu, C., & Rudd, C. E. Decreased progenitor TCF1+ T-cells correlate with COVID-19 disease severity. differentiation, 43, 45. https://doi.org/10.1038/s42003-024-05922-2La Pietra, A., Bianchi, A. R., Capriello, T., Mobilio, T., Guagliardi, A., De Maio, A., & Ferrandino, I. (2024). Regeneration of zebrafish retina following toxic injury. Environmental Toxicology and Pharmacology, 112, 104582. https://doi.org/10.1016/j.etap.2024.104582La Pietra, A., Imperatore, R., Coccia, E., Mobilio, T., Ferrandino, I., & Paolucci, M. (2024). Comparative Study of Condensed and Hydrolysable Tannins during the Early Stages of Zebrafish Development. International Journal of Molecular Sciences, 25(13), 7063. https://doi.org/10.3390/ijms25137063Lettieri, A., Oleari, R., van den Munkhof, M. H., van Battum, E. Y., Verhagen, M. G., Tacconi, C., ... & Cariboni, A. (2023). SEMA6A drives GnRH neuron-dependent puberty onset by tuning median eminence vascular permeability. Nature Communications, 14(1), 8097. https://doi.org/10.1038/s41467-023-43820-zLi, C., Lu, Y., Wang, J., Liu, B., Szeto, I. M. Y., Zhang, W., ... & Sun, Y. (2024). Immunoregulation of bovine lactoferrin together with osteopontin promotes immune system development and maturation. Food & Function, 15(2), 866-880. https://doi.org/10.1039/D3FO03515HLi, F., Liu, J., Liu, C., Liu, Z., Peng, X., Huang, Y., ... & Wu, D. (2024). Cyclic peptides discriminate BCL-2 and its clinical mutants from BCL-XL by engaging a single-residue discrepancy. Nature Communications, 15(1), 1476. https://doi.org/10.1038/s41467-024-45848-1Li, G., Qian, Y., Chen, Y., Cao, M., Yang, X., Kong, D., ... & Liu, Y. (2022). Wip1 contributes to the adaptation of HepG2 human liver cancer cells to stress hormone-induced DNA damage. Oncology Letters, 25(1), 31. https://doi.org/10.3892/ol.2022.13617Li, L., Yang, Y., Wang, Z., Xu, C., Huang, J., & Li, G. (2021). Prognostic role of METTL1 in glioma. Cancer cell international, 21, 1-20. https://doi.org/10.1186/s12935-021-02346-4Li, L., Yao, Y., Cao, L., Le, Y., Li, X., Wang, X., ... & Gong, P. (2025). RAGE-mediated intestinal pro-inflammatory responses triggered by Giardia duodenalis. Acta Tropica, 107529. https://doi.org/10.1016/j.actatropica.2025.107529Li, M., Liu, G., Jin, X., Guo, H., Setrerrahmane, S., Xu, X., ... & Xu, H. (2022). Micropeptide MIAC inhibits the tumor progression by interacting with AQP2 and inhibiting EREG/EGFR signaling in renal cell carcinoma. Molecular cancer, 21(1), 181. https://doi.org/10.1186/s12943-022-01654-1Li, R., Xiong, W., Li, B., Li, Y., Fang, B., Wang, X., & Ren, F. (2023). Plasmalogen Improves Memory Function by Regulating Neurogenesis in a Mouse Model of Alzheimer’s Diseases. International Journal of Molecular Sciences, 24(15), 12234. https://doi.org/10.3390/ijms241512234Li, T., Niu, Z., Yu, T., Li, J., Lu, X., Huang, M., ... & Shan, H. (2023). Nucleosome assembly protein 1 like 1 (NAP1L1) promotes cardiac fibrosis by inhibiting YAP1 ubiquitination and degradation. MedComm, 4(5), e348. https://doi.org/10.1002/mco2.348Li, X., Chen, C., Chen, Y., Jiang, K., Zhao, X., Zhang, F., & Li, Y. (2024). Oridonin ameliorates ocular surface inflammatory responses by inhibiting the NLRP3/caspase-1/GSDMD pyroptosis pathway in dry eye. Experimental Eye Research, 109955. https://doi.org/10.1016/j.exer.2024.109955Li, Y., Du, X., Hu, Y., Wang, D., Duan, L., Zhang, H., ... & Cui, Y. (2024). Iron-laden macrophage-mediated paracrine profibrotic signaling induces lung fibroblast activation. American Journal of Physiology-Cell Physiology, 327(4), C979-C993. https://doi.org/10.1152/ajpcell.00675.2023Li, Y., Lin, Y., Tang, Y., Jiang, M., Chen, X., Chen, H., ... & Wang, C. (2024). MAZ-mediated up-regulation of BCKDK reprograms glucose metabolism and promotes growth by regulating glucose-6-phosphate dehydrogenase stability in triple-negative breast cancer. Cell Death & Disease, 15(7), 516. https://doi.org/10.1038/s41419-024-06835-yLi, Y., Tang, L., Wang, F., Gao, C., Yang, Q., Luo, L., ... & Qi, M. (2024). Hub genes identification and validation of ferroptosis in SARS-CoV-2 induced ARDS: perspective from transcriptome analysis. Frontiers in Immunology, 15, 1407924. https://doi.org/10.3389/fimmu.2024.1407924Li, Y., Wang, Z., Bai, L. L., Li, Y. Z., Jiang, Y. J., Xu, T. L., ... & Zhao, X. (2024). Positive Intervention of Distinct Peptides in Clostridioides difficile Infection in a Mouse Model. Communications Biology, 7(1), 1172. https://doi.org/10.1038/s42003-024-06850-xLi, Z., Liu, S., Zhu, T., An, X., Wei, X., Zhang, J., ... & Wan, X. (2022). The loss-function of the male sterile gene ZmMs33/ZmGPAT6 results in severely oxidative stress and metabolic disorder in maize anthers. Cells, 11(15), 2318. https://doi.org/10.3390/cells11152318Liang, X., Miao, Y., Tong, X., Chen, J., Liu, H., He, Z., ... & Hu, Z. (2024). Dental pulp mesenchymal stem cell-derived exosomes inhibit neuroinflammation and microglial pyroptosis in subarachnoid hemorrhage via the miRNA-197-3p/FOXO3 axis. Journal of Nanobiotechnology, 22(1), 426. https://doi.org/10.1186/s12951-024-02708-wLiao, X., Cai, D., Liu, J., Hu, H., You, R., Pan, Z., ... & Huang, H. (2023). Deletion of Mettl3 in mesenchymal stem cells promotes acute myeloid leukemia resistance to chemotherapy. Cell Death & Disease, 14(12), 796. https://doi.org/10.1038/s41419-023-06325-7Licini, C., Morroni, G., Lucarini, G., Vitto, V. A. M., Orlando, F., Missiroli, S., ... & Marchi, S. (2024). ER-mitochondria association negatively affects wound healing by regulating NLRP3 activation. Cell Death & Disease, 15(6), 407. https://doi.org/10.1038/s41419-024-06765-9Liu, C., Gong, J. S., Su, C., Li, H., Qin, J., Xu, Z. H., & Shi, J. S. (2023). Increasing gene dosage and chaperones co-expression facilitate the efficient dextranase expression in Pichia pastoris. LWT, 181, 114753. https://doi.org/10.1016/j.lwt.2023.114753Liu, F., You, F., Yang, L., Wang, S., & Xie, D. (2024). Metformin improves diabetic neuropathy by reducing inflammation through up-regulating the expression of miR-146a and suppressing oxidative stress. Journal of Diabetes and its Complications, 38(6), 108737. https://doi.org/10.1016/j.jdiacomp.2024.108737Liu, G., Hou, Y., Jin, X., Zhang, Y., Sun, C., Huang, C., ... & Jiang, X. (2024). PI3K/HSCB axis facilitates FOG1 nuclear translocation to promote erythropoiesis and megakaryopoiesis. Elife, 13, RP95815. https://doi.org/10.7554/eLife.95815.3Liu, L., Gao, J., Tang, Y., Guo, G., & Gan, H. (2023). Increased expression of the P2Y12 receptor is involved in the failure of autogenous arteriovenous fistula caused by stenosis. Renal Failure, 45(2), 2278314. https://doi.org/10.1080/0886022X.2023.2278314Liu, P., Li, H., Xu, H., Gong, J., Jiang, M., Xu, Z., & Shi, J. (2023). Aggravated hepatic fibrosis induced by phenylalanine and tyrosine was ameliorated by chitooligosaccharides supplementation. Iscience, 26(10). https://doi.org/10.1016/j.isci.2023.107754Liu, S., Nam, H. S., Zeng, Z., Deng, X., Pashaei, E., Zang, Y., ... & Wan, J. (2024). CDHu40: a novel marker gene set of neuroendocrine prostate cancer. Briefings in bioinformatics, 25(6), bbae471. https://doi.org/10.1093/bib/bbae471Liu, X., Xu, J., Zhang, M., Wang, H., Guo, X., Zhao, M., ... & Guo, Y. (2024). RABV induces biphasic actin cytoskeletal rearrangement through Rac1 activity modulation. Journal of Virology, e00606-24. https://doi.org/10.1128/jvi.00606-24Liu, Y. J., Chen, X. F., Zhou, L. P., Rao, F., Zhang, D. Y., & Wang, Y. H. (2022). A nerve conduit filled with Wnt5a‐loaded fibrin hydrogels promotes peripheral nerve regeneration. CNS neuroscience & therapeutics, 28(1), 145-157. https://doi.org/10.1111/cns.13752Liu, Y., Wu, J., Najem, H., Lin, Y., Pang, L., Khan, F., ... & Chen, P. (2024). Dual targeting macrophages and microglia is a therapeutic vulnerability in models of PTEN-deficient glioblastoma. The Journal of Clinical Investigation, 134(22). https://doi.org/10.1172/JCI178628Lopes-Paciencia, S., Bourdeau, V., Rowell, M. C., Amirimehr, D., Guillon, J., Kalegari, P., ... & Ferbeyre, G. (2024). A senescence restriction point acting on chromatin integrates oncogenic signals. Cell Reports, 43(4). https://doi.org/10.1016/j.celrep.2024.114044Lu, B., Meng, R., Wang, Y., Xiong, W., Ma, Y., Gao, P., ... & Yuan, X. (2024). Distinctive physiological and molecular responses of foxtail millet and maize to nicosulfuron. Frontiers in Plant Science, 14, 1308584. https://doi.org/10.3389/fpls.2023.1308584Lu, J. C. L. (2022). Transcriptional regulation of type 2 innate lymphoid cells by interleukin-7 receptor signaling (Doctoral dissertation, University of British Columbia). https://dx.doi.org/10.14288/1.0417452Lu, S., Hernan, R., Marcogliese, P. C., Huang, Y., Gertler, T. S., Akcaboy, M., ... & Bellen, H. J. (2022). Loss-of-function variants in TIAM1 are associated with developmental delay, intellectual disability, and seizures. The American Journal of Human Genetics, 109(4), 571-586. https://doi.org/10.1016/j.ajhg.2022.01.020Lu, Z., Zhang, J., Wang, H., Zhang, K., Gu, Z., Xu, Y., ... & Zhou, C. (2024). Rewiring of a KNOXI regulatory network mediated by UFO underlies the compound leaf development in Medicago truncatula. Nature Communications, 15(1), 2988. https://doi.org/10.1038/s41467-024-47362-wLuo, N., Cheng, W., Zhou, Y., Gu, B., Zhao, Z., & Zhao, Y. (2021). Screening candidate genes regulating placental development from trophoblast transcriptome at early pregnancy in Dazu Black goats (Capra hircus). Animals, 11(7), 2132. https://doi.org/10.3390/ani11072132Luo, N., Xiong, K., Wang, Y., Cen, C., Liu, Q., Wang, F., ... & Hou, X. (2024). Coridius chinensis Extract Promotes Leydig Cell Endocrine Function Rescuing Testosterone Deficiency and Sexual Behavior Insufficiency in Manganese‐Exposed Rat. Andrologia, 2024(1), 6629144. https://doi.org/10.1155/2024/6629144Luo, N., Zhou, Y., Chen, X., Zhao, Y., & Hu, Y. (2024). Screening the optimal housekeeping genes (HKGs) of placenta tissues by RNA-sequence and qRT-PCR throughout gestation in goat (Capra Hircus). Gene, 895, 147966. https://doi.org/10.1016/j.gene.2023.147966Ma, C., Zheng, S., Yang, S., Wu, J., Sun, X., Chen, Y., ... & Chen, R. (2025). OsCYCBL1 and OsHTR702 positively regulate rice tolerance to cold stress. International Journal of Biological Macromolecules, 287, 138642. https://doi.org/10.1016/j.ijbiomac.2024.138642Ma, M., Zhang, X., Zheng, Y., Lu, S., Pan, X., Mao, X., ... & Bellen, H. J. (2023). The fly homolog of SUPT16H, a gene associated with neurodevelopmental disorders, is required in a cell-autonomous fashion for cell survival. Human Molecular Genetics, 32(6), 984-997. https://doi.org/10.1093/hmg/ddac259Ma, P., Zhang, Y., Yin, Y., Wang, S., Chen, S., Liang, X., ... & Deng, H. (2024). Gut microbiota metabolite tyramine ameliorates high-fat diet-induced insulin resistance via increased Ca2+ signaling. The EMBO Journal, 43(16), 3466-3493. https://doi.org/10.1038/s44318-024-00162-wMang, Y., Zhang, S., Zhao, J., Ran, J., Zhao, Y., Li, L., ... & Bao, F. (2022). Characteristics of pre-sensitization-related acute antibody-mediated rejection in a rat model of orthotopic liver transplantation. Annals of Translational Medicine, 10(19). https://doi.org/10.21037/atm-22-4311Méant, A., Moussa, O., Lebeau, B., Gonçalves, C., Richard, V. R., Cai, F., ... & Witcher, M. (2024). Combined Inhibition of MNK Signaling and BET Proteins Reveals TGM2 as a Novel Vulnerability in Melanoma. Journal of Investigative Dermatology. https://doi.org/10.1016/j.jid.2024.07.037Meng, J., Lv, Q., Sui, A., Xu, D., Zou, T., Song, M., ... & Wang, X. (2022). Hyperuricemia induces lipid disturbances by upregulating the CXCL-13 pathway. American Journal of Physiology-Gastrointestinal and Liver Physiology, 322(2), G256-G267. https://doi.org/10.1152/ajpgi.00285.2021Meng, R., Li, Z., Kang, X., Zhang, Y., Wang, Y., Ma, Y., ... & Wang, J. (2024). High Overexpression of SiAAP9 Leads to Growth Inhibition and Protein Ectopic Localization in Transgenic Arabidopsis. International Journal of Molecular Sciences, 25(11), 5840. https://doi.org/10.3390/ijms25115840Miao, Y., Liang, X., Chen, J., Liu, H., He, Z., Qin, Y., ... & Zhang, R. (2024). Transfer of miR-877–3p via extracellular vesicles derived from dental pulp stem cells attenuates neuronal apoptosis and facilitates early neurological functional recovery after cerebral ischemia–reperfusion injury through the Bclaf1/P53 signaling pathway. Pharmacological Research, 206, 107266. https://doi.org/10.1016/j.phrs.2024.107266
-
Milholland, K. L., AbdelKhalek, A., Baker, K. M., Hoda, S., DeMarco, A. G., Naughton, N. H., ... & Hall, M. C. (2022). Reduced Cdc14 phosphatase activity impairs septation, hyphal differentiation and pathogenesis and causes echinocandin hypersensitivity in Candida albicans. bioRxiv, 2022-09. https://doi.org/10.1101/2022.09.29.510203
Milholland, K. L., AbdelKhalek, A., Baker, K. M., Hoda, S., DeMarco, A. G., Naughton, N. H., ... & Hall, M. C. (2023). Cdc14 phosphatase contributes to cell wall integrity and pathogenesis in Candida albicans. Frontiers in Microbiology, 14, 1129155. https://doi.org/10.3389/fmicb.2023.1129155
Mohammed, S. K., Rasheed, M. N., & Asker, B. A. (2024). The expression of genes TLR2 and TLR10 in the gastric tissue of patients with gastroduodenal disorders caused by Helicobacter pylori. Baghdad Science Journal, 21(8), 2500-2500. https://doi.org/10.21123/bsj.2023.8962
Muñoz-Villagrán, C., Acevedo-Arbunic, J., Härtig, E., Sievers, S., Zühlke, D., Issotta, F., ... & Levicán, G. (2024). The Thioredoxin Fold Protein (TFP2) from Extreme Acidophilic Leptospirillum sp. CF-1 Is a Chaperedoxin-like Protein That Prevents the Aggregation of Proteins under Oxidative Stress. International Journal of Molecular Sciences, 25(13), 6905. https://doi.org/10.3390/ijms25136905
Nassir, N., Tambi, R., Bankapur, A., Al Heialy, S., Karuvantevida, N., Khansaheb, H. H., ... & Uddin, M. (2021). Single-cell transcriptome identifies FCGR3B upregulated subtype of alveolar macrophages in patients with critical COVID-19. Iscience, 24(9). https://doi.org/10.1016/j.isci.2021.103030
Nassir, N., Tambi, R., Bankapur, A., Karuvantevida, N., Khansaheb, H. H., Zehra, B., ... & Uddin, M. (2022). Analyzing single cell transcriptome data from severe COVID-19 patients. STAR protocols, 3(2), 101379. https://doi.org/10.1016/j.xpro.2022.101379
Ninfole, E. (2022). Involvement of autophagy in cholestatic liver diseases. https://hdl.handle.net/11566/298981
Niu, X., Zhang, F., Ping, L., Wang, Y., Zhang, B., Wang, J., & Chen, X. (2023). vwa1 knockout in zebrafish causes abnormal craniofacial chondrogenesis by regulating FGF pathway. Genes, 14(4), 838. https://doi.org/10.3390/genes14040838
Palma, D., Oliva, V., Montanares, M., Gil-Durán, C., Travisany, D., Chávez, R., & Vaca, I. (2024). Expanding the Toolbox for Genetic Manipulation in Pseudogymnoascus: RNAi-Mediated Silencing and CRISPR/Cas9-Mediated Disruption of a Polyketide Synthase Gene Involved in Red Pigment Production in P. verrucosus. Journal of Fungi, 10(2), 157. https://doi.org/10.3390/jof10020157
Pang, L., Dunterman, M., Guo, S., Khan, F., Liu, Y., Taefi, E., ... & Chen, P. (2023). Kunitz-type protease inhibitor TFPI2 remodels stemness and immunosuppressive tumor microenvironment in glioblastoma. Nature Immunology, 24(10), 1654-1670. https://doi.org/10.1038/s41590-023-01605-y
Panja, S., Truica, M. I., Yu, C. Y., Saggurthi, V., Craige, M. W., Whitehead, K., ... & Mitrofanova, A. (2024). Mechanism-centric regulatory network identifies NME2 and MYC programs as markers of Enzalutamide resistance in CRPC. Nature communications, 15(1), 352. https://doi.org/10.1038/s41467-024-44686-5
Parra, M., Aldabaldetrecu, M., Arce, P., Soto-Aguilera, S., Vargas, R., Guerrero, J., ... & Modak, B. (2024). [Cu (NN1) 2] ClO4, a Copper (I) Complex as an Antimicrobial Agent for the Treatment of Piscirickettsiosis in Atlantic Salmon. International Journal of Molecular Sciences, 25(7), 3700. https://doi.org/10.3390/ijms25073700
Parra, M., Aldabaldetrecu, M., Arce, P., Soto-Aguilera, S., Vargas, R., Guerrero, J., ... & Modak, B. (2024). Oral administration of a new copper (I) complex with coumarin as ligand: modulation of the immune response and the composition of the intestinal microbiota in Onchorhynchus mykiss. Frontiers in Chemistry, 12, 1338614. https://doi.org/10.3389/fchem.2024.1338614
Pernet, E., Downey, J., Vinh, D. C., Powell, W. S., & Divangahi, M. (2019). Leukotriene B4–type I interferon axis regulates macrophage-mediated disease tolerance to influenza infection. Nature microbiology, 4(8), 1389-1400. https://doi.org/10.1038/s41564-019-0444-3
Pernet, E., Sun, S., Sarden, N., Gona, S., Nguyen, A., Khan, N., ... & Divangahi, M. (2023). Neonatal imprinting of alveolar macrophages via neutrophil-derived 12-HETE. Nature, 614(7948), 530-538. https://doi.org/10.1038/s41586-022-05660-7
Pham, H. M., & Do, T. T. (2023). Detection and assessment of risk factors associated with Newcastle disease virus infection in birds in backyard poultry in Laichau province of Vietnam. Avian Pathology, 52(2), 144-152. https://doi.org/10.1080/03079457.2022.2160697
Pu, Y., Cheng, R., Zhang, Q., Huang, T., Lu, C., Tang, Z., ... & Liu, Y. (2023). Role of soluble epoxide hydrolase in the abnormal activation of fibroblast-like synoviocytes from patients with rheumatoid arthritis. Clinical Immunology, 257, 109850. https://doi.org/10.1016/j.clim.2023.109850
Pu, Y., He, Y., Zhao, X., Zhang, Q., Wen, J., Hashimoto, K., & Liu, Y. (2022). Depression-like behaviors in mouse model of Sjögren's syndrome: A role of gut–microbiota–brain axis. Pharmacology Biochemistry and Behavior, 219, 173448. https://doi.org/10.1016/j.pbb.2022.173448
Qi, L., Duan, B. W., Wang, H., Liu, Y. J., Han, H., Han, M. M., ... & Li, L. (2024). Reactive Oxygen Species‐Responsive Nanoparticles Toward Extracellular Matrix Normalization for Pancreatic Fibrosis Regression. Advanced Science, 11(19), 2401254. https://doi.org/10.1002/advs.202401254
Qiu, L., Hu, M., Qin, X., Song, R., Sun, Y., & Wang, X. (2024). Intracellular regulation limits the response of intestinal ferroportin to Iron Status in Suckling rats. Molecular Nutrition & Food Research, 68(6), 2300617. https://doi.org/10.1002/mnfr.202300617
Quan, R., Shi, C., Fang, B., Sun, Y., Qu, T., Wang, X., ... & Li, Y. (2024). Age-Dependent Inflammatory Microenvironment Mediates Alveolar Regeneration. International Journal of Molecular Sciences, 25(6), 3476. https://doi.org/10.3390/ijms25063476
Quan, R., Shi, C., Sun, Y., Zhang, C., Bi, R., Zhang, Y., ... & Li, Y. (2024). PAI-1 Derived from Alveolar Type 2 Cells Drives Aging-Associated Pulmonary Fibrosis. Engineering, 42, 74-87. https://doi.org/10.1016/j.eng.2024.08.014
Rahmy, S., Mishra, S. J., Murphy, S., Blagg, B. S., & Lu, X. (2022). Hsp90β inhibition upregulates interferon response and enhances immune checkpoint blockade therapy in murine tumors. Frontiers in Immunology, 13, 1005045. https://doi.org/10.3389/fimmu.2022.1005045
Ren, Y., Kong, M., Sun, H., Zhao, B., Wu, H., Chen, Z., ... & Zhang, Q. (2024). Genome-wide identification, characterization and expression profiling of TLR family genes in Chromileptes altivelis. Fish & Shellfish Immunology, 109720. https://doi.org/10.1016/j.fsi.2024.109720
Rendine, M., Cocci, P., de Vivo, L., Bellesi, M., & Palermo, F. A. (2024). Effects of Chronic Sleep Restriction on Transcriptional Sirtuin 1 Signaling Regulation in Male Mice White Adipose Tissue. Current Issues in Molecular Biology, 46(3), 2144-2154. https://doi.org/10.3390/cimb46030138
Reyes González, J. I. (2023). Expresión de los factores de transcripción SOX5 y Twist y su relación con quimioresistencia en modelos in vitro de cáncer gástrico. https://doi.org/10.58011/f9z2-z534
Robichon, L., Bar, C., Marian, A., Lehmann, L., Renault, S., Kabashi, E., ... & Nabbout, R. (2024). kcnb1 loss-of-function in zebrafish causes neurodevelopmental and epileptic disorders associated with GABA dysregulation. bioRxiv, 2024-07. https://doi.org/10.1101/2024.07.03.601913
Rozza, N., Grunbaum, A. M., Kremer, R., & Milhalcioiu, C. The identification of a SARs-CoV2 S2 protein derived peptide with super-antigen-like stimulatory properties on T-cells. https://doi.org/10.1038/s42003-024-07350-8
Rui-Qi, Z., Tian, P., Hong-Cen, W., Zi-Fan, W., Xue-Feng, W., Chang-Yong, Z., & Shi-Min, F. (2024). Identification and analysis of bZIP family genes in Citrus sinensis and the role of CsbZIP24 in response to Huanglongbing. Scientia Horticulturae, 336, 113436. https://doi.org/10.1016/j.scienta.2024.113436
Ruiting, L. U. O., Huang, Y., Ruimin, B. A. I., Meng, L. I. U., Liang, S. U. N., Xiaofei, W. A. N. G., & Zheng, Y. (2024). ATP Citrate Lyase is a General Tumour Biomarker and Contributes to the Development of Cutaneous Squamous Cell Carcinoma. Acta Dermato-venereologica, 104. https://doi.org/10.2340/actadv.v104.23805
Saha, D., Gregor, J. B., Hoda, S., Eastman, K. E., Navarrete, M., Wisecaver, J. H., & Briggs, S. D. (2024). Candida glabrata maintains two Hap1 homologs, Zcf27 and Zcf4, for distinct roles in ergosterol gene regulation to mediate sterol homeostasis under azole and hypoxic conditions. bioRxiv. https://doi.org/10.1101/2024.06.20.599910
Sander, N. H., Soni, S., Wilkinson, C. M., Khiabani, E., Dyck, J. R., & Colbourne, F. (2024). Exogenous ketone therapy does not protect brain tissue after moderate-sized intracerebral hemorrhage despite signs of early neurological benefit. Plos one, 19(12), e0311778. https://doi.org/10.1371/journal.pone.0311778
Sang, X., Wang, Q., Ning, Y., Wang, H., Zhang, R., Li, Y., ... & Ren, F. (2023). Age-related mucus barrier dysfunction in mice is related to the changes in muc2 mucin in the colon. Nutrients, 15(8), 1830. https://doi.org/10.3390/nu15081830
Schweiger, M. W., Amoozgar, Z., Repiton, P., Morris, R., Maksoud, S., Hla, M., ... & Tannous, B. A. (2024). Glioblastoma extracellular vesicles modulate immune PD-L1 expression in accessory macrophages upon radiotherapy. Iscience, 27(2). https://doi.org/10.1016/j.isci.2024.108807
Shen, Z., Gao, Y., Sun, X., Chen, M., Cen, C., Wang, M., ... & Gao, F. (2024). Inactivation of JNK signalling results in polarity loss and cell senescence of Sertoli cell. Cell Proliferation, e13760. https://doi.org/10.1111/cpr.13760
Shi, X., Wang, Z., Guo, M., Wang, Y., Bi, Z., Li, D., ... & Liu, J. (2023). PRP coating on different modified surfaces promoting the osteointegration of polyetheretherketone implant. Frontiers in Bioengineering and Biotechnology, 11, 1283526. https://doi.org/10.3389/fbioe.2023.1283526
Shi, Y., Chen, Z., Shen, M., Li, Q., Wang, S., Jiang, J., & Zeng, W. (2024). Identification and Functional Verification of the Glycosyltransferase Gene Family Involved in Flavonoid Synthesis in Rubus chingii Hu. Plants, 13(10), 1390. https://doi.org/10.3390/plants13101390
Silva-Sanzana, C., Zavala, D., Moraga, F., Herrera-Vásquez, A., & Blanco-Herrera, F. (2022). Oligogalacturonides enhance resistance against aphids through pattern-triggered immunity and activation of salicylic acid signaling. International Journal of Molecular Sciences, 23(17), 9753. https://doi.org/10.3390/ijms23179753
Song, B., Du, J., & Vleeshouwers, V. (Eds.). (2023). Characterization of major traits and identification of functional genes for potato. Frontiers Media SA. https://doi.org/10.3389/978-2-8325-2668-2
Song, Y., Hao, L., Wang, X., Wang, X., Cong, P., Li, Z., ... & Xu, J. (2024). Temperature effects on plasmalogen profile and quality characteristics in Pacific oyster (Crassostrea gigas) during depuration. Food Research International, 186, 114356. https://doi.org/10.1016/j.foodres.2024.114356
Su, T., Li, Z., Zhang, Y., Xu, J., & Xu, B. (2024). Regulatory Mechanisms of Strigolactones on the Development of Lateral Branches in Cucumber. Journal of the American Society for Horticultural Science, 149(3), 162-168. https://doi.org/10.21273/JASHS05384-24
Sudo, H., & Kubo, A. (2021). The aneugenicity of ketone bodies in colon epithelial cells is mediated by microtubule hyperacetylation and is blocked by resveratrol. International Journal of Molecular Sciences, 22(17), 9397. https://doi.org/10.3390/ijms22179397
Sun, Y., Jin, C., Wu, S., Yin, C., Chen, J., Bao, Z., ... & Hu, J. (2024). Identification and functional characterization of IGFBP genes in Leopard Coral Grouper (Plectropomus leopardus): Insights into growth regulation and stress response. Water Biology and Security, 100338. https://doi.org/10.1016/j.watbs.2024.100338
Tang, W., Yan, H., Chen, X., Pu, Y., Qi, X., Dong, L., & Su, C. (2024). hUCMSC-derived extracellular vesicles relieve cisplatin-induced granulosa cell apoptosis in mice by transferring anti-apoptotic miRNAs. The Journal of Biomedical Research. https://dx.doi.org/10.7555/JBR.37.20230310
Tang, X., Morris, A. J., Deken, M. A., & Brindley, D. N. (2023). Autotaxin inhibition with IOA-289 decreases breast tumor growth in mice whereas knockout of autotaxin in adipocytes does not. Cancers, 15(11), 2937. https://doi.org/10.3390/cancers15112937
Tao, J., Zhu, S., Liao, X., Wang, Y., Zhou, N., Li, Z., ... & Liu, R. (2022). DLP-based bioprinting of void-forming hydrogels for enhanced stem-cell-mediated bone regeneration. Materials today Bio, 17, 100487. https://doi.org/10.1016/j.mtbio.2022.100487
Tao, J., Zhu, S., Zhou, N., Wang, Y., Wan, H., Zhang, L., ... & Liu, R. (2022). Nanoparticle‐stabilized emulsion bioink for digital light processing based 3D bioprinting of porous tissue constructs. Advanced Healthcare Materials, 11(12), 2102810. https://doi.org/10.1002/adhm.202102810
Tiwari, A., Hashemiaghdam, A., Laramie, M. A., Maschi, D., Haddad, T., Stunault, M. I., ... & Ashrafi, G. (2023). Sirtuin3 ensures the metabolic plasticity of neurotransmission during glucose deprivation. Journal of Cell Biology, 223(1), e202305048. https://doi.org/10.1083/jcb.202305048
Tiwari, A., Myeong, J., Hashemiaghdam, A., Stunault, M. I., Zhang, H., Niu, X., ... & Ashrafi, G. (2024). Mitochondrial pyruvate transport regulates presynaptic metabolism and neurotransmission. Science Advances, 10(46), eadp7423. https://doi.org/10.1101/2024.03.20.586011
Tram, J., Marty, L., Mourouvin, C., Abrantes, M., Jaafari, I., Césaire, R., ... & Peloponese, J. M. J. (2024). The Oncoprotein Fra-2 Drives the Activation of Human Endogenous Retrovirus Env Expression in Adult T-Cell Leukemia/Lymphoma (ATLL) Patients. Cells, 13(18), 1517. https://doi.org/10.3390/cells13181517
Tu, T. H., Bennani, F. E., Masroori, N., Liu, C., Nemati, A., Rozza, N., ... & Rudd, C. E. (2025). The identification of a SARs-CoV2 S2 protein derived peptide with super-antigen-like stimulatory properties on T-cells. Communications Biology, 8(1), 14. https://doi.org/10.5281/zenodo.14036310
Tu, T. H., Grunbaum, A., Santinon, F., Kazanova, A., Rozza, N., Kremer, R., ... & Rudd, C. E. (2024). Decreased progenitor TCF1+ T-cells correlate with COVID-19 disease severity. Communications biology, 7(1), 526. https://doi.org/10.1038/s42003-024-05922-2
Wang, C., He, Y., Fang, X., Zhang, D., Huang, J., Zhao, S., ... & Li, G. (2024). METTL1-Modulated LSM14A Facilitates Proliferation and Migration in Glioblastoma via the Stabilization of DDX5. iScience. https://doi.org/10.1016/j.isci.2024.110225
Wang, C., Zhang, X., Wang, X., Zhai, Y., Li, M., Pan, J., ... & Zhou, J. (2022). Genetic deletion of hspa8 leads to selective tissue malformations in zebrafish embryonic development. Journal of cell science, 135(21), jcs259734. https://doi.org/10.1242/jcs.259734
Wang, D., Zhang, T., Qiu, L., & Zhao, C. (2024). The Potential of the Probiotic Isolate Lactobacillus plantarum SS18‐50 to Prevent Colitis in Mice. Food Science & Nutrition. https://doi.org/10.1002/fsn3.4657
Wang, H. H., Sun, Y. N., Qu, T. Q., Sang, X. Q., Zhou, L. M., Li, Y. X., & Ren, F. Z. (2022). Nobiletin prevents D-Galactose-Induced C2C12 cell aging by improving mitochondrial function. International journal of molecular sciences, 23(19), 11963. https://doi.org/10.3390/ijms231911963
Wang, H. H., Zhang, Y., Qu, T. Q., Sang, X. Q., Li, Y. X., Ren, F. Z., ... & Sun, Y. N. (2023). Nobiletin improves D-galactose-induced aging mice skeletal muscle atrophy by regulating protein homeostasis. Nutrients, 15(8), 1801. https://doi.org/10.3390/nu15081801
Wang, H. Q., Liu, Y., Li, D., Liu, J. Y., Jiang, Y., He, Y., ... & Sun, H. X. (2023). Maternal and embryonic signals cause functional differentiation of luminal epithelial cells and receptivity establishment. Developmental Cell, 58(21), 2376-2392. https://doi.org/10.1016/j.devcel.2023.08.004
Wang, H., Song, S., Gao, S., Yu, Q., Zhang, H., Cui, X., ... & Qi, T. (2024). The NLR immune receptor ADR1 and lipase-like proteins EDS1 and PAD4 mediate stomatal immunity in Nicotiana benthamiana and Arabidopsis. The Plant Cell, 36(2), 427-446. https://doi.org/10.1093/plcell/koad270
Wang, H., Tan, J., Cui, X., Bai, Y., Gao, S., Staskawicz, B., ... & Qi, T. (2024). Switch of TIR signaling by a Ca2+ sensor activates ADR1 recognition of pRib-AMP-EDS1-PAD4 for stomatal immunity. bioRxiv, 2024-10. https://doi.org/10.1101/2024.10.29.620780
Wang, J., Peng, Z., Guo, J., Wang, Y., Wang, S., Jiang, H., ... & Liu, Z. (2023). CXCL10 Recruitment of γδ T Cells into the Hypoxic Bone Marrow Environment Leads to IL17 Expression and Multiple Myeloma Progression. Cancer Immunology Research, 11(10), 1384-1399. https://doi.org/10.1158/2326-6066.CIR-23-0088
Wang, M., Li, J., Liu, B., Shen, Z., Chen, M., Cui, X., ... & Zhao, H. (2025). TRAPPC2l Participates in Male Germ Cell Development by Regulating Cell Division. Cell Proliferation, e13810. https://doi.org/10.1111/cpr.13810
Wang, W., Shi, J., Zhang, Y., Qiao, Y., Zuo, W., Wang, Z., ... & Wei, P. (2023). Genetic and pathogenic characterizations of a naturally occurring reassortant and homologous recombinant strain of the classical infectious bursal disease virus re-emerging in chickens in southern China. Frontiers in Microbiology, 14, 1293072. https://doi.org/10.3389/fmicb.2023.1293072
Wang, X., Shen, T., Lian, J., Deng, K., Qu, C., Li, E., ... & Li, X. (2023). Resveratrol reduces ROS-induced ferroptosis by activating SIRT3 and compensating the GSH/GPX4 pathway. Molecular Medicine, 29(1), 137. https://doi.org/10.1186/s10020-023-00730-6
Wang, X., Sun, Y., Luan, C., Yang, S., Wang, K., Zhang, X., ... & Zhang, W. (2024). Effect of hydrogen‐rich saline on melanopsin after acute blue light‐induced retinal damage in rats. Photochemistry and Photobiology. https://doi.org/10.1111/php.13952
Wang, Y., Zhou, X., Zhu, S., Wei, X., Zhou, N., Liao, X., ... & Liu, R. (2022). Cryoprinting of nanoparticle-enhanced injectable hydrogel with shape-memory properties. Materials & Design, 223, 111120. https://doi.org/10.1016/j.matdes.2022.111120
Wang, Z., Zhang, X., Liu, C., Duncan, S., Hang, R., Sun, J., ... & Cao, X. (2024). AtPRMT3-RPS2B promotes ribosome biogenesis and coordinates growth and cold adaptation trade-off. Nature Communications, 15(1), 8693. https://doi.org/10.1038/s41467-024-52945-8
Wei, X., Zheng, T., Zhu, S., Liao, X., Zhou, N., Wang, Y., ... & Tao, J. (2024). Fe3O4‐nanoparticle‐functionalized 3D‐printed injectable microgel. Journal of Applied Polymer Science, 141(6), e54913. https://doi.org/10.1002/app.54913
Wojtas, M. N., Diaz-González, M., Stavtseva, N., Shoam, Y., Verma, P., Buberman, A., ... & Knafo, S. (2024). Interplay between hippocampal TACR3 and systemic testosterone in regulating anxiety-associated synaptic plasticity. Molecular Psychiatry, 29(3), 686-703. https://doi.org/10.1038/s41380-023-02361-z
Wu, F., Liu, Y., Zhang, M., Yuan, X., Ji, T., Jin, Y., ... & Fang, B. (2024). Effects of 1-oleate-2-palmitate-3-linoleate glycerol supplementation on the small intestinal development and gut microbial composition of neonatal mice. Food Research International, 195, 114993. https://doi.org/10.1016/j.foodres.2024.114993
Wu, J., Chen, Q., Wang, Y., Wang, R., Chen, Q., Wang, Y., ... & Chen, K. (2024). LINC01977 promotes colorectal cancer growth and metastasis by enhancing aerobic glycolysis via the ERK/c-Myc axis. Journal of Gastrointestinal Oncology, 15(1), 271. https://doi.org/10.21037/jgo-24-52
Wu, J., Muhammad, T., Hu, J., Zhao, L., Wang, J., & Liu, X. (2025). SpWRKY71 is required for drought tolerance in tomato. New Zealand Journal of Crop and Horticultural Science, 1-20. https://doi.org/10.1080/01140671.2025.2455050
Wu, K., Duan, X., Zhu, Z., Sang, Z., Zhang, Y., Li, H., ... & Ma, L. (2021). Transcriptomic analysis reveals the positive role of abscisic acid in endodormancy maintenance of leaf buds of Magnolia wufengensis. Frontiers in Plant Science, 12, 742504. https://doi.org/10.3389/fpls.2021.742504
Wu, W. B., Xu, Y. Y., Cheng, W. W., Yuan, B., Zhao, J. R., Wang, Y. L., & Zhang, H. J. (2017). Decreased PGF may contribute to trophoblast dysfunction in fetal growth restriction. Reproduction, 154(3), 319-329. https://doi.org/10.1530/REP-17-0253
Wu, X., Lin, T., Zhou, X., Zhang, W., Liu, S., Qiu, H., ... & Tian, Z. (2024). Potato E 3 ubiquitin ligase S t RFP 1 positively regulates late blight resistance by degrading sugar transporters S t SWEET10 c and S t SWEET 11. New Phytologist. https://doi.org/10.1111/nph.19848
Wu, X., Zhou, X., Lin, T., Zhang, Z., Wu, X., Zhang, Y., ... & Tian, Z. (2024). Accumulation of dually targeted StGPT1 in chloroplasts mediated by StRFP1, an E3 ubiquitin ligase, enhances plant immunity. Horticulture Research, 11(11), uhae241. https://doi.org/10.1093/hr/uhae241
Xia, Y., Feng, Y., Jiang, L., Heng, Y., Li, X., & Ma, C. (2024). The Effects of Metabolites from Three Vaginal Bacteria on the Syndecan-1 of Cervical Epithelial Cells. Heliyon, e33426. https://doi.org/10.1016/j.heliyon.2024.e33426
Xie, X. D., Zhou, Y., Sun, Y. B., Yi, S. L., Zhao, Y., Chen, Q., ... & Hu, T. J. (2023). RNA-Seq and 16S rRNA reveals that Tian–Dong–Tang–Gan Powder alleviates environmental stress-induced decline in immune and antioxidant function and gut microbiota dysbiosis in Litopenaeus vannami. Antioxidants, 12(6), 1262. https://doi.org/10.3390/antiox12061262
Xiong, W., Li, R., Li, B., Wang, X., Wang, H., Sun, Y., ... & Ren, F. (2023). Nobiletin mitigates D-galactose-induced memory impairment via improving hippocampal neurogenesis in mice. Nutrients, 15(9), 2228. https://doi.org/10.3390/nu15092228
Xu, B., Huang, J. P., Peng, G., Cao, W., Liu, Z., Chen, Y., ... & Huang, S. X. (2024). Total biosynthesis of the medicinal triterpenoid saponin astragalosides. Nature Plants, 1-12. https://doi.org/10.1101/gr.276196.121
Xu, L., Huang, X., Chen, Z., Yang, M., & Deng, J. (2024). Eosinophil peroxidase promotes bronchial epithelial cells to secrete asthma-related factors and induces the early stage of airway remodeling. Clinical Immunology, 263, 110228. https://doi.org/10.1016/j.clim.2024.110228
Xu, P., Zhuo, W., Zhang, P., Chen, Y., Du, Y., Li, Y., & Wang, Y. (2025). Cyclin G1 Regulates the Alveolarization in Models of Bronchopulmonary Dysplasia by Inhibiting AT2 Cell Proliferation. Biomolecules, 15(1), 101. https://doi.org/10.3390/biom15010101
Xu, W., Rustenhoven, J., Nelson, C. A., Dykstra, T., Ferreiro, A., Papadopoulos, Z., ... & Kipnis, J. (2023). A novel immune modulator IM33 mediates a glia-gut-neuronal axis that controls lifespan. Neuron, 111(20), 3244-3254. https://doi.org/10.1016/j.neuron.2023.07.010
Xuan, W., Hsu, W. H., Khan, F., Dunterman, M., Pang, L., Wainwright, D. A., ... & Chen, P. (2022). Circadian regulator CLOCK drives immunosuppression in glioblastoma. Cancer immunology research, 10(6), 770-784. https://doi.org/10.1158/2326-6066.CIR-21-0559
Yan, B., Hooper, D. C., Yuan, Z., Wang, C., Chen, Y., & Lu, B. (2023). Autoantibodies drive heart damage caused by concomitant radiation and PD-1 Blockade. Cancer Immunology Research, 11(4), 546-555. https://doi.org/10.1158/2326-6066.CIR-21-0839
Yan, K., Jin, C., Men, Y., Chen, Y., Li, Z., Cai, W., ... & Qi, J. (2024). The role of plin2a in glycolysis regulation in Sertoli cells and its cascading impact on spermatogonial development in black rockfish (Sebastes schlegelii). Water Biology and Security, 100335. https://doi.org/10.1016/j.watbs.2024.100335
Yang, C., Guo, L., Du, J., Zhang, Q., & Zhang, L. (2024). SPINK1 Overexpression Correlates with Hepatocellular Carcinoma Treatment Resistance Revealed by Single Cell RNA-Sequencing and Spatial Transcriptomics. Biomolecules, 14(3), 265. https://doi.org/10.3390/biom14030265
Yang, J., Ji, J., Sun, Y. Q., Li, S. Z., Jiang, Z. K., Dong, Y. F., & Sun, X. L. (2022). Nuclear localization of platelet-activating factor receptor accounts for microglial phagocytosis in ischemic stroke. https://doi.org/10.21203/rs.3.rs-1732120/v1
Yang, R., Li, L., Hou, Y., Li, Y., Zhang, J., Yang, N., ... & Shan, H. (2023). Long non-coding RNA KCND1 protects hearts from hypertrophy by targeting YBX1. Cell Death & Disease, 14(5), 344. https://doi.org/10.1038/s41419-023-05852-7
Yang, S., Tian, M., Dai, Y., Feng, S., Wang, Y., Chhangani, D., ... & Johnson, A. (2022). Infection and chronic disease activate a brain-muscle signaling axis that regulates muscle performance. bioRxiv, 2020-12. https://doi.org/10.1101/2020.12.20.423533
Yang, Y., Cai, Q., Luo, L., Sun, Z., & Li, L. (2024). Genome-Wide Analysis of C-Repeat Binding Factor Gene Family in Capsicum baccatum and Functional Exploration in Low-Temperature Response. Plants, 13(4), 549. https://doi.org/10.3390/plants13040549
Yang, Y., Cai, Q., Wang, X., Yang, Y., Li, L., Sun, Z., & Li, W. (2025). Transcriptomic and Metabolomic Analyses Reveal Differences in Flavonoid Synthesis During Fruit Development of Capsicum frutescens pericarp. Agriculture, 15(2), 222. https://doi.org/10.3390/agriculture15020222
Yang, Y., Cai, Q., Wang, Y., Li, L., & Sun, Z. (2024). Genome-Wide Exploration of the WD40 Gene Family in Eggplant (Solanum melongena L.) and Analysis of Its Function in Fruit Color Formation. Agronomy, 14(3), 521. https://doi.org/10.3390/agronomy14030521
Yang, Y., Cai, Q., Yang, Y., Wang, X., Li, L., Sun, Z., & Li, W. (2024). Transcriptomics and Metabolomics Reveal Biosynthetic Pathways and Regulatory Mechanisms of Phenylpropanes in Different Ploidy of Capsicum frutescens. Plants, 13(23), 3393. https://doi.org/10.3390/plants13233393
Yi, S., Cai, Q., Yang, Y., Shen, H., Sun, Z., & Li, L. (2024). Identification and Functional Characterization of the SaMYB113 Gene in Solanum aculeatissimum. Plants, 13(11), 1570. https://doi.org/10.3390/plants13111570
Yin, Y., Ma, P., Wang, S., Zhang, Y., Han, R., Huo, C., ... & Deng, H. (2022). The CRTC-CREB axis functions as a transcriptional sensor to protect against proteotoxic stress in Drosophila. Cell Death & Disease, 13(8), 688. https://doi.org/10.1038/s41419-022-05122-y
Yu, P., Xu, T., Ma, W., Fang, X., Bao, Y., Xu, C., ... & Li, G. (2024). PRMT6-mediated transcriptional activation of ythdf2 promotes glioblastoma migration, invasion, and emt via the wnt–β-catenin pathway. Journal of Experimental & Clinical Cancer Research, 43(1), 116. https://doi.org/10.1186/s13046-024-03038-3
Yu, S., Wang, X., Li, Z., Jin, D., Yu, M., Li, J., ... & Ren, F. (2024). Solobacterium moorei promotes the progression of adenomatous polyps by causing inflammation and disrupting the intestinal barrier. Journal of Translational Medicine, 22(1), 169. https://doi.org/10.1186/s12967-024-04977-3
Zhan, J., Wu, S., Zhao, X., & Jing, J. (2021). A novel DNA damage repair-related gene signature for predicting glioma prognosis. International Journal of General Medicine, 10083-10101. https://doi.org/10.2147/IJGM.S343839
Zhang, F., Song, W., Yang, R., Jin, C., Xie, Y., Shen, Y., ... & He, Y. (2024). Semen promotes oocyte development in Sebastes schlegelii elucidating ovarian development dynamics in live-bearing fish. Iscience, 27(3). https://doi.org/10.1016/j.isci.2024.109193
Zhang, L., Smyth, D., Al-Khalaf, M., Blet, A., Du, Q., Bernick, J., ... & Liu, P. P. (2022). Insulin-like growth factor-binding protein-7 (IGFBP7) links senescence to heart failure. Nature Cardiovascular Research, 1(12), 1195-1214. https://doi.org/10.1038/s44161-022-00181-y
Zhang, M., Luo, X., He, W., Zhang, M., Peng, Z., Deng, H., & Xing, J. (2024). OsJAZ4 Fine-Tunes Rice Blast Resistance and Yield Traits. Plants, 13(3), 348. https://doi.org/10.3390/plants13030348
Zhang, S., Wang, X., He, J., Zhang, S., Zhao, T., Fu, S., & Zhou, C. (2023). A Sec-dependent effector, CLIBASIA_04425, contributes to virulence in ‘Candidatus Liberibater asiaticus’. Frontiers in Plant Science, 14, 1224736. https://doi.org/10.3389/fpls.2023.1224736
Zhang, Y., Diao, H. T., Leng, M. Y., Wu, Y. Z., Huang, B. Y., Li, X., ... & Guo, J. (2025). YTHDF3-mediated FLCN/cPLA2 axis improves cardiac fibrosis via suppressing lysosomal function. Acta Pharmacologica Sinica, 1-13. https://doi.org/10.1038/s41401-024-01425-2
Zhang, Y., Liu, Y., Teng, Z., Wang, Z., Zhu, P., Wang, Z., ... & Liu, X. (2023). Human umbilical cord mesenchymal stem cells (hUC-MSCs) alleviate paclitaxel-induced spermatogenesis defects and maintain male fertility. Biological Research, 56(1), 47. https://doi.org/10.1186/s40659-023-00459-w
Zhang, Y., Zheng, Z., Gao, J., Bao, X., Zhang, W., Liu, L., ... & Li, Y. (2024). Rare-Earth Metal-Based Nanosystems for Facilitating Neural Stem Cell Differentiation into Neurons and Enhancing Axonal Stability. ACS Applied Nano Materials, 7(14), 16154-16161. https://doi.org/10.1021/acsanm.4c02057
Zhang, Z., Zhang, Y., Qiu, Y., Mo, W., & Yang, Z. (2021). Human/eukaryotic ribosomal protein L14 (RPL14/eL14) overexpression represses proliferation, migration, invasion and EMT process in nasopharyngeal carcinoma. Bioengineered, 12(1), 2175-2186. https://doi.org/10.1080/21655979.2021.1932225
Zhao, G., Zhang, X., Meng, L., Dong, K., Shang, S., Jiang, T., ... & Gao, H. (2024). Single-cell RNA-sequencing reveals a unique landscape of the tumor microenvironment in obesity-associated breast cancer. Oncogene, 43(45), 3277-3290. https://doi.org/10.1038/s41388-024-03161-7
Zhao, J., Gong, F., Yang, Q., Yang, R., Yan, Z., Xi, Z., ... & Liu, X. (2024). Exercise in ozone-polluted air evokes pathological cardiac hypertrophy via up-regulation of nuclear lncRNA EYA4-au1 and recruiting Med11 to activating EYA4/p27kip1/CK2α/HDAC2 cascade. Ecotoxicology and Environmental Safety, 287, 117264. https://doi.org/10.1016/j.ecoenv.2024.117264
Zhao, L., Wang, B., Yang, T., Li, N., Yang, H., Wang, J., & Yan, H. (2024). Investigation of the regulation of drought tolerance by the SlHVA22l gene in tomato. Chinese Bulletin of Botany, 59(4), 558-573. https://doi.org/10.11983/CBB23129
Zhao, S. Q., Chen, M. J., Chen, F., Gao, Z. F., Li, X. P., Hu, L. Y., ... & Song, Z. W. (2025). ENTPD8 Overexpression Enhances Anti-PD-L1 Therapy in Hepatocellular Carcinoma via miR-214-5p Inhibition. iScience. https://doi.org/10.1016/j.isci.2025.111819
Zhao, X., Chen, Y., Li, R., Men, Y., Yan, K., Li, Z., ... & Qi, J. (2024). Immune Rejection Mediated by prf1 and gzmb Affects the Colonization of Fat Greenling (Hexagrammos otakii) Spermatogonia in Heterotransplantation. International Journal of Molecular Sciences, 25(10), 5157. https://doi.org/10.3390/ijms25105157
Zhao, X., Zhu, Z., Sang, Z., Ma, L., Yin, Q., & Jia, Z. (2024). Physiological and Transcriptomic Analyses Demonstrate the Ca2+-Mediated Alleviation of Salt Stress in Magnolia wufengensis. Plants, 13(17), 2418. https://doi.org/10.3390/plants13172418
Zhao, Y., Xie, Q., Yang, Q., Cui, J., Tan, W., Zhang, D., ... & Yan, M. (2024). Genome-wide identification and evolutionary analysis of the NRAMP gene family in the AC genomes of Brassica species. BMC Plant Biology, 24(1), 311. https://doi.org/10.1186/s12870-024-04981-1
Zheng, Z., Yuan, D., Shen, C., Zhang, Z., Ye, J., & Zhu, L. (2023). Identification of potential diagnostic biomarkers of atherosclerosis based on bioinformatics strategy. BMC Medical Genomics, 16(1), 100. https://doi.org/10.1186/s12920-023-01531-w
Zhong, X., Zeng, H., Zhou, Z., Su, Y., Cheng, H., Hou, Y., ... & Ding, J. (2023). Structural mechanisms for regulation of GSDMB pore-forming activity. Nature, 616(7957), 598-605. https://doi.org/10.1038/s41586-023-05872-5
Zhou, J. X., Su, X. M., Zheng, S. Y., Wu, C. J., Su, Y. N., Jiang, Z., ... & He, X. J. (2022). The Arabidopsis NuA4 histone acetyltransferase complex is required for chlorophyll biosynthesis and photosynthesis. Journal of integrative plant biology, 64(4), 901-914. https://doi.org/10.1111/jipb.13227
Zhou, J., Guo, M., Yang, G., Cui, X., Hu, J., Lin, T., ... & Wang, Y. (2024). Chromatin landscape dynamics during reprogramming towards human naïve and primed pluripotency reveals the divergent function of PRDM1 isoforms. Cell Death Discovery, 10(1), 474. https://doi.org/10.1038/s41420-024-02230-w
Zhou, J., Liu, X., Dong, Q., Li, J., Niu, W., & Liu, T. (2024). Extracellular vesicle-bound VEGF in oral squamous cell carcinoma and its role in resistance to Bevacizumab Therapy. Cancer Cell International, 24(1), 296. https://doi.org/10.1186/s12935-024-03476-1
Zhou, Y., Li, X., Zhang, X., Li, M., Luo, N., & Zhao, Y. (2023). Screening of candidate housekeeping genes in uterus caruncle by RNA-Sequence and qPCR analyses in different stages of goat (Capra hircus). Animals, 13(12), 1897. https://doi.org/10.3390/ani13121897
Zi, T., YaNan, L., ZeLin, W., YuSheng, Z., MeiNa, X., Peng, Z., ... & XueXia, L. (2023). Melatonin alleviates oxidative damage in mouse spermatogenesis and sperm quality parameters induced by exposure to Bisphenol A. Ecotoxicology and environmental safety, 253, 114709. https://doi.org/10.1016/j.ecoenv.2023.114709
Zu, X., Luo, L., Wang, Z., Gong, J., Yang, C., Wang, Y., ... & Cao, X. (2023). A mitochondrial pentatricopeptide repeat protein enhances cold tolerance by modulating mitochondrial superoxide in rice. Nature Communications, 14(1), 6789. https://doi.org/10.1038/s41467-023-42269-4
김상섭. (2023). 초파리의 Pss 유전자 기능 상실 돌연변이에 의한 미토콘드리아 이상 및 근감소증 연구 (Doctoral dissertation, 서울대학교 대학원). https://hdl.handle.net/10371/204018